All published articles of this journal are available on ScienceDirect.
COVID-19 Origins: A Mixed-Methods Meta-Analysis of Scientific Consensus and Political Narratives
Abstract
Introduction
Since late 2019, the origin of COVID-19 has been a topic of debate among scientists and politicians, particularly between the US and China. The lab leak hypothesis, which involves the escape of the virus from the WIV laboratory in Wuhan, China, and the natural origin hypothesis, possibly through an intermediate host. These hypotheses have been discussed since the onset of the pandemic without a definitive conclusion. However, a prevailing view within the scientific community suggests a specific origin, potentially unbiased by political influences. This study aims to investigate the direction of scientific consensus on the origin of matter and to discuss the effects and impacts of politicization.
Method
To achieve this, a mixed-method meta-analysis involving a content-based qualitative and quantitative synthesis was conducted. Forty-eight studies were selected using the PRISMA model and were synthesized in MASQDA. Textual analyses included text processing, TF-IDF weighting, sentiment scoring using AFINN lexicon, and similarity metrics (Jaccard, Levenshtein) to map inter-document relationships and key evidentiary terms. A sensitivity analysis was performed to verify the robustness of our AFINN-based continuous sentiment measurements by comparing them to Bing, Loughran, Syuzhet, NRC, and VADER lexicons. The preregistration of this protocol occurred on PROSPERO with an ID: 1055566.
Results
According to the results derived from the study, many scientists support the natural origin of COVID-19, and the sentiment around this hypothesis is positive (0.398), indicating a more optimistic and affirmative language compared to the lab-leak hypothesis, with a negative average sentiment score of -0.124, suggesting that discourse around this theory is comparatively more negative. However, these findings were consistent across multiple sentiment analysis tools (Bing, Loughran, Syuzhet, NRC, and VADER), confirming that the lab-leak narrative tends to be discussed more negatively, which provides a robustness of the main AFINN results, while Natural Origin narratives tend toward more positive sentiment. Therefore, several key factors contribute to the scientific preference for the natural origin hypothesis, including: (1) No record of genetic evidence of laboratory engineering has been found. (2) No pre-existing virus matching SARS-CoV-2 was known to be held in any lab (3) The furin cleavage site (FCS) is naturally occurring and experimental attempts to generate an FCS in bat coronaviruses failed, suggesting natural evolution, (4)) early cases linked to animal exposure, not labs, (5) historical precedent for natural zoonotic spillover: SARS-CoV-1 (2003) and MERS-CoV (2012), (6) lack of credible evidence for lab involvement: no scientific publication, leaked document, or whistleblower testimony.
Discussion
The analysis reveals a strong scientific inclination toward the natural origin hypothesis of COVID-19, indicating a positive sentiment score reflecting more confident and supportive language in the literature compared to the lab-leak hypothesis. Although the lab-leak hypothesis is often discussed, it is usually described in more negative and uncertain terms. This revealed that both hypotheses are part of the academic conversation; the natural origin theory has more substantial evidence and more supportive discourse. However, this scientific debate has become highly politicized, especially between the U.S. and China. This political friction has muddled public understanding and threatens to erode trust in science. In particular, the lab-leak narrative has been frequently promoted outside of scientific circles on platforms with political motivations, fueling polarized public arguments.
Conclusion
This study illustrates the impact of political polarization on scientific communication and perception, and how public debate can change in relation to scientific consensus. To preserve the integrity of science, investigations into viral origins must be transparent, cooperative, and free from geopolitical influence. This commitment is essential to ensure preparedness for future pandemics and maintain public trust in science.
1. INTRODUCTION
On 11th February 2020, a novel respiratory syndrome coronavirus 2 (SARS-CoV-2), as named by the ICTV, was associated with the disease officially named COVID-19, resulting in a pandemic that emerged as one of the most significant global crises of the 21st century [1, 2]. Since its first reported cases in Wuhan, China, in late 2019, the COVID-19 virus has claimed many lives, resulting in a high rate of death, economic crises, and disruption in daily life around the world [3, 2]. Many scientists have focused on understanding the mode of transmission [4], therapy [5, 6], and prevention associated with the spread of the virus. However, one significant question remained unanswered: What is the true origin of SARS-CoV-2?
The controversial debate over the origin of COVID-19 has been centered on two mutually exclusive primary hypotheses: (1) the natural origin hypothesis, which suggests the zoonotic origin of the virus, and (2) a laboratory-related origin hypothesis, which involves the accidental escape of the virus from the Wuhan Institute of Virology (WIV) research facility in China [2, 7]. These two hypotheses are not merely academic or scientific; they touch upon sensitive geopolitical dynamics, national accountability, and the integrity of global scientific collaboration. In 2021, the World Health Organization (WHO) embarked on an investigative mission which stated that a lab leak was “extremely unlikely,” but acknowledged that more research was needed. Meanwhile, governments, intelligence agencies, and scientists have made different and sometimes conflicting claims, further introducing more complexity. Public discourse has also been shaped by misinformation, ideological biases, and politically motivated narratives. Indeed, what began as a virological inquiry has gradually escalated into a battleground of media sensationalism, political posturing, and social polarization.
Understanding the origin of SARS-CoV-2 is crucial not only for establishing historical facts surrounding the origin of the virus but also for improving future pandemic preparedness, strengthening laboratory safety rules, and guiding public health policies [8, 9]. It is equally important to examine how origin narratives are constructed, communicated, and politicized [7, 10]. These narratives significantly influence public opinion, scientific research, and government decisions. They also affect international relationships, especially between major global powers such as the United States and China, whose relationship has been tested by accusations and counter-accusations regarding COVID-19’s genesis [10, 7].
Therefore, the reasons for studying the origin of COVID-19 should not be limited to focusing only on where the virus first appeared. The study should also investigate how scientific evidence, politics, and the media work together to shape global understanding. This study adopts an interdisciplinary approach, using both qualitative and quantitative methods to analyze the main narratives about the origin of COVID-19 and how they evolve. This layered analysis can offer critical insights into the intersection of science, politics, and society in the 21st century. In light of the complex and multifaceted nature of the COVID-19 origin debate, this study seeks to explore the issue from an interdisciplinary perspective, employing both qualitative and quantitative methods to examine the dominant narratives surrounding the origin of COVID-19 and how they have evolved over time. The scientific evidence supports each central hypothesis (zoonotic i. lab-based), and how has this evidence been represented in academic and public discourse, and to what extent has the investigation into COVID-19's origins been influenced by political, institutional, and geopolitical factors? Finally, it shows how politicization has affected scientific communication, research integrity, and public trust.
2. INTERMEDIATE HOST
Scientifically, a host is an animal or plant on or in which a parasite or commensal organism lives [11]. According to the Biology Online Dictionary, there are five types of hosts, which are: primary, secondary, paratenic, accidental, and reservoir [12]. Out of these, the secondary host (also known as the intermediate host) in disease transmission is significant. The definition of secondary host comes from its ability to serve as a passage (i.e., intermediate) for pathogen transmission. An intermediate host is an organism that temporarily harbors a pathogen-such as a virus-allowing it to replicate or mutate before the pathogen is transmitted to its final or primary host, often a human [13, 14, 15]. In zoonotic diseases (infections that jump from animals to humans), the intermediate host serves as a biological bridge between the natural reservoir and humans [16, 17]. The primary role of an intermediate host is to facilitate viral adaptation and amplification [18, 19]. Within the intermediate host, the virus may increase in concentration and undergo genetic changes or recombination, which can enhance its ability to bind to human receptors and cause infection [20, 21]. While animals are overwhelmingly the known intermediate hosts in zoonotic transmission, theoretically, humans can also act as intermediate hosts in anthroponotic or reverse-zoonotic events-where a virus originates in animals [17], infects humans, and is then passed on to other animals or other humans with evolutionary shifts [22, 23, 24, 25]. However, in classical zoonotic emergence (such as SARS, MERS, and potentially COVID-19), intermediate hosts have always been animals, as reported by numerous historical data including viruses that transmit directly to humans such as HIV [26], Marburg virus [27], Rabies virus [28], Hantavirus [29], Monkeypox virus [30], and Lassa virus [31], as well as viruses requiring intermediate hosts like Dengue virus [32], Yellow Fever virus [33], Zika virus [34], Influenza A [35], Nipah virus [36], Hendra virus [37], Ebola virus [38], MERS-CoV [39], SARS-CoV [40], and potentially SARS-CoV-2 via pangolins [41] (Fig. 1).
3. METHODOLOGY
3.1. Research Design
To ensure the study captures the full complexity of the COVID-19 origins debate, we employed a mixed-method meta-analysis instead of a single-method study because of the contested nature of the COVID-19 origin debate. Hence, the mixed-method meta-analysis allows the extraction and synthesis of qualitative and quantitative data from findings across multiple sources and disciplines [42]. This synthesis is crucial, since no single study or perspective has definitively resolved the origin debate. At the same time, the Meta-analysis allows the combination of diverse perspectives, identifies areas of consensus or disagreement, and evaluates the methodological strengths and weaknesses across the existing research [43]. The quantitative analysis of this study involves systematic data aggregation and statistical synthesis from peer-reviewed literature and public databases [42]. Metrics such as the frequency of specific claims about the virus's origin, shifts in the scientific consensus over time, patterns of international co-authorship, and the referenced impact of relevant studies. Qualitatively, thematic content analysis was performed to explain why certain narratives gain traction or how political and ideological forces influence the scientific process and public discourse [42, 44].
3.2. Search Strategy
The search strategy we employed in this study was focused on trustworthy sources of information, including peer-reviewed articles, government reports, and official statements indexed in widely recognized databases, such as PubMed, Scopus, Web of Science, and Embase, supplemented by Google Scholar for broader coverage and the World Health Organization (WHO) website for authoritative global reports. These sources were selected because they are trusted worldwide for accurate and wide-ranging research, especially in health and science, which helped to include only high-quality, relevant studies in the review. These databases were queried using a combination of predefined keywords and Boolean operators, including: (“COVID-19” OR “SARS-CoV-2”) AND (“origin” OR “source” OR “spillover” OR “zoonotic” OR “lab leak” OR “Wuhan” OR “animal origin”). Additional keyword phrases were used to enhance search sensitivity, such as “COVID-19 origin,” “SARS-CoV-2 origin,” “origin of COVID-19,” and “origin of SARS-CoV-2. Finally, Studies were selected based on predefined inclusion and exclusion criteria listed below (Table 1).
| Criteria | Inclusion | Exclusion |
|---|---|---|
| Study Type | Peer-reviewed articles, systematic reviews, meta-analyses, narrative reviews, and institutional or governmental reports | Editorials, commentaries, unpublished articles, and letters |
| Language | English | Non-English publications |
| Publication Date | 2019 – July 2025 | Published before Jan 2020 |
| Focus | Studies investigating COVID-19 origin hypotheses (natural spillover, lab leak) | Studies do not focus on the origin of COVID-19 |
| Accessibility | Open | Closed |
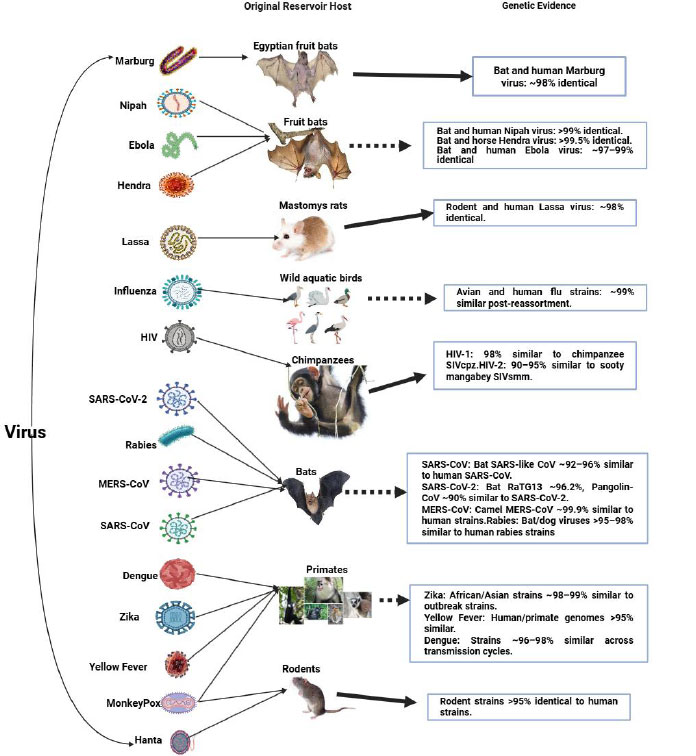
Genetic similarity between animal reservoir viruses and human virus strains. The thick dotted lines indicate transmission through an intermediate host, while the thick lines indicate direct transmission to humans.
The review protocol was registered on PROSPERO (ID: 1055566) and followed PRISMA guidelines [Supplementary 2]. The initial search with the keywords returned a total of 142,314 records, including 5,286 from PubMed, 53 from SAGE Journals, approximately 124,000 from Google Scholar (top 500 screened by relevance), and 50 from Elsevier (Fig. 2).
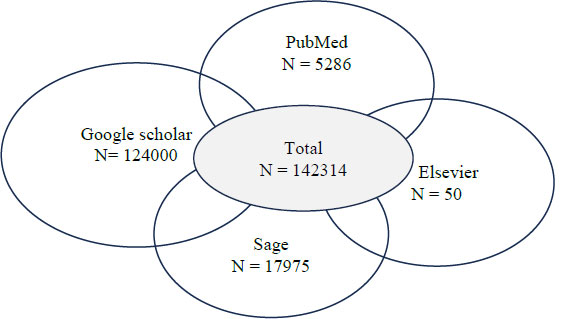
Number of studies retrieved across databases.
Following the initial search, 2,319 duplicates were removed, and 145,000 studies were subsequently screened. A total of 144,100 records were excluded, after which 900 studies were sought for retrieval, and 50 were not retrieved. On applying the aforementioned inclusion and exclusion criteria, 628 studies were excluded. Two hundred twenty-two studies were included in the review, and 48 studies were synthesized (Fig. 3) [Supplementary 1 contains the complete list of synthesized documents].
3.3. Qualitative Analysis
The thematic analysis was done to extract meaningful patterns, statements, and narratives from textual materials. Each of the identified documents was renamed with a unique document ID (D1, D2, D3, … Dn) for consistency and easy identification during the coding [Supplementary 3 contains the anonymized source documents with their corresponding IDs]. These IDs match the study ID in the paper metadata [Supplementary 4], collected on an Ms. Excel sheet which contains: reference, title, source, abstract, etc., The document (i.e., articles, reports, etc.) was carefully read multiple times to gain a general sense of recurring ideas and themes, after which the text was broken into segments and manually coded for significant content. Codes were clustered into thematic categories based on conceptual similarity and frequency of appearance. The coding framework was done inductively and deductively-initial codes emerged from the literature, while others were based on theoretical frameworks such as framing theory and science-politics interface theory [45]. Themes were categorized into: Natural origin theory, lab-leak origin theory, scientific consensus, politicization of the COVID-19 origin, why lab-leak fails, prior event, Unknown origin, recommendation and future direction, and impact of global policy as shown in Table S1 (Supplementary File 5). The coding was performed using the Maxqda qualitative analytic pro 2020 tool version 20.3.1 [46].
3.4. Quantitative Analysis
The extracted data were standardized by tokenization, stop word removal, and lemmatization for comparison across sources. Descriptive statistics, such as frequencies, proportions, means, and medians, were done for temporal trend (annual publication/citation metrics) and source distribution. Hypothesis support was quantified through segment-level frequency analysis, comparing “Lab-Leak” and “Natural Origin” prevalence across subgroups (author fields, institutions, countries) using a percentage breakdown. Textual analyses included TF-IDF weight keyword extraction, keyness metrics (likelihood ratio) [47] to contrast lexical and affective features between hypotheses. The AFINN lexicon [48] was used to capture the intensity (dislike and hate are considered negative words with different intensities) without relying on informal linguistic markers (., exclamation points) that are absent in this dataset to measure the sentiment around the original discussion. Each token contributed equally, and scores were kept continuous without thresholds. Similarity metrics Jaccard [49, 50, 51] and Levenshtein [52, 53] were used to map inter-document textual relationships, while targeted phrase extraction identified evidentiary keywords (., “zoonotic,” “spillover”) [see Supplementary 6 & Supplementary 7 for similarity metric results]. Data were processed and analyzed using R statistical software.
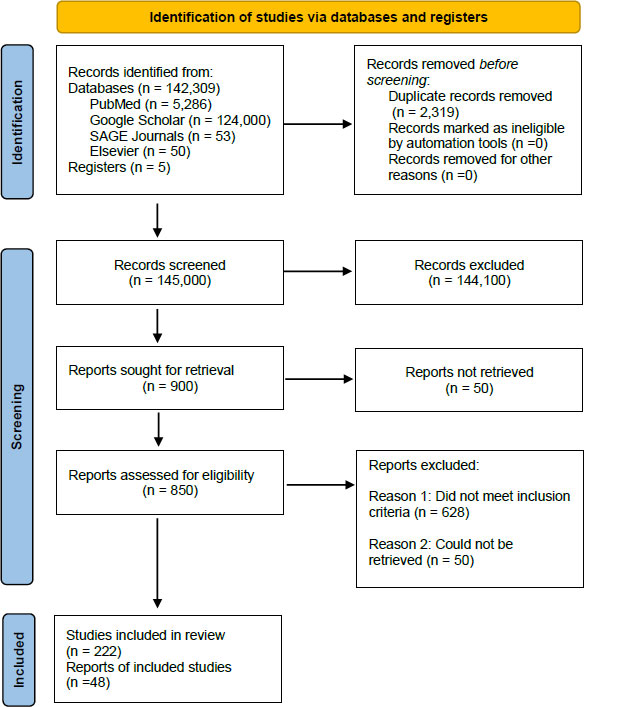
The Preferred reporting items for systematic reviews and meta-analyses (PRISMA) model indicating the study selection process.
3.5. Sensitive and Robust Analysis
We complemented AFINN with alternative lexicons (Bing, Loughran, NRC, Syuzhet, and VADER) to conduct sensitive checks, ensuring that results are not an artifact of one sentiment scheme. To ensure robustness, we evaluated alternative weighting schemes. First, we normalized sentiment scores by document length. Second, we applied tf–idf weighting to reduce the influence of widespread, domain-generic terms. Finally, we compared mean scores to a polarity balance measure (positive-to-negative ratio).
3.6. Validity and Reliability
Construct validity was ensured by clearly defining each thematic category and cross-verifying data sources. Internal validity was supported by triangulating findings from multiple sources and applying inter-coder agreement in qualitative coding. Similarly, external validity was cautiously considered, with an acknowledgment that findings may be more representative of English-speaking scientists. For qualitative data, inter-coder reliability was maintained by employing two researchers for the thematic coding process, who conducted a manual thematic analysis to avoid a semantic equivalence between a contradictory word (., not originate naturally and originate naturally). To ensure reliability and consistency, a third-party reviewer verified the coded dataset and facilitated discussions where discrepancies arose. Any disagreements between coders were resolved through consensus discussion involving the third-party reviewer, ensuring that the final codes accurately reflected the content. For quantitative datasets, reliability was ensured by validating data collection procedures, running duplicate queries for verification, and using standardized meta-analytical methods.
4. RESULT AND DISCUSSION
Out of the 48 selected documents (Fig. 4), the majority of the papers were published in 2020(20[41.67%]) with 12,608(94.6%, 788.0 (±1273.8),80.5 (IQR: 0–873.5) citations. This likely reflects the urgency and global focus on COVID-19 during the early phase of the pandemic. In contrast, papers from 2021 (15[31.25%]) and 2022 (6[12.50%]), 2023(5[10.42%]), 2024(2[4.17%], and 2025(1[2.08%]) experienced a significant decline in publication and citations [see Table S1 for distribution of included papers and citation metrics by year of publication (Supplementary 8)].
4.1. The Origin Debate: Zoonotic Hypothesis and Lab-origin Hypothesis
A total of 699 discrete statements were identified that explicitly supported either the Lab Leak or the Natural Origin Hypothesis regarding the origin of COVID-19. There were 111(15.9%) statements that supported the Lab Leak Hypothesis and 588(84.1%) statements that supported the Natural Origin Hypothesis (Fig. 5).
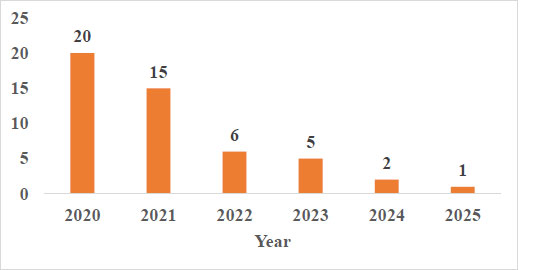
Distribution of included papers by year of publication.
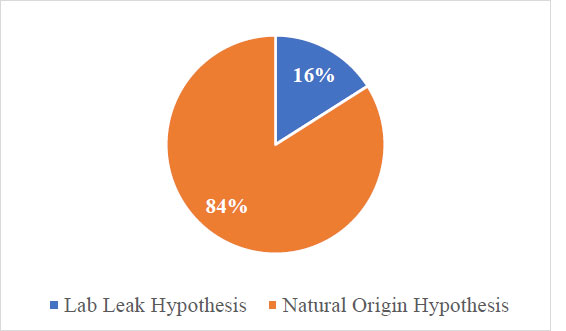
Distribution of total discrete statements supporting COVID-19 origin hypotheses.
The analysis of the TF-IDF scores for words associated with the Natural Origin Hypothesis and the Lab Leak Hypothesis reveals distinct thematic focuses in the language used across both narratives (Fig. 6) [with complete TF-IDF weighted keyword tables provided in Supplementary 9]. For the Natural Origin Hypothesis, the top term was “wildlife, indicating that this word was both frequent and uniquely representative of this hypothesis. In contrast, the Lab Leak Hypothesis contains language suggestive of institutional processes and investigative discourse. [Supplementary 9].
As shown in Fig. (7), the keyness analysis finds which words are most strongly linked to the Natural Origin i lab leak narrative in the COVID-19 origin debate. The word “bat” appears to have the highest linked word with a G2 value (47.98, p < 0.0), followed by pangolin. This revealed that these animals are central to the discourse of the zoonotic transmission hypothesis. In contrast, the word “laboratory” is associated with the lab leak idea, followed by the USA representing a key word in the theory (P < 0.00) [see Supplementary 10 for full keyness metrics].
The sentiment analysis of the two narratives, the Lab Leak Hypothesis and the Natural Origin Hypothesis, reveals a notable emotional divergence in how each is discussed in the literature (Fig. 8). The Natural Origin Hypothesis is associated with a positive average sentiment score of 0.398, indicating a more optimistic and affirmative language. However, the Lab Leak Hypothesis shows a negative average sentiment score of -0.124, suggesting that discourse around this theory is comparatively more negative. This suggests that the languages used for this hypothesis are characterized by skepticism, criticism, controversy, and suspicion. The tone reflects defensive or accusatory framing, particularly in politicized or non-peer-reviewed sources. The sensitive analysis reaffirmed this pattern [see Tables S2-S4 and Figures S1-S3 (Supplementary File 8)]: alternative lexicons (Bing, Loughran, Syuzhet, VADER) consistently produced negative sentiment scores for Lab Leak, with only NRC yielding a slight positive signal. For Natural Origin, NRC and VADER confirmed positive sentiment, though Bing, Loughran, and Syuzhet returned weaker or negative values. Taken together, these analyses confirm the robustness of the main finding: Lab Leak narratives are framed more negatively, whereas Natural Origin narratives tend toward more positive sentiment.
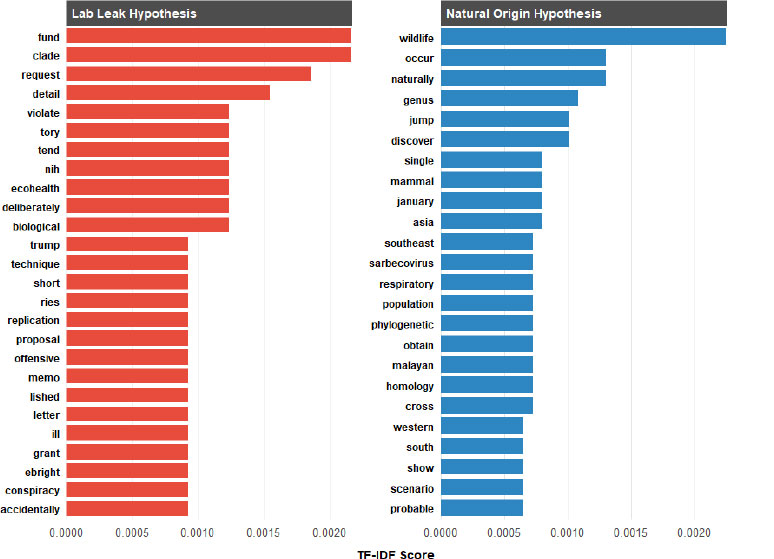
The 20 Distinctive terms ranked by TF-IDF score for each COVID-19 origin hypothesis.
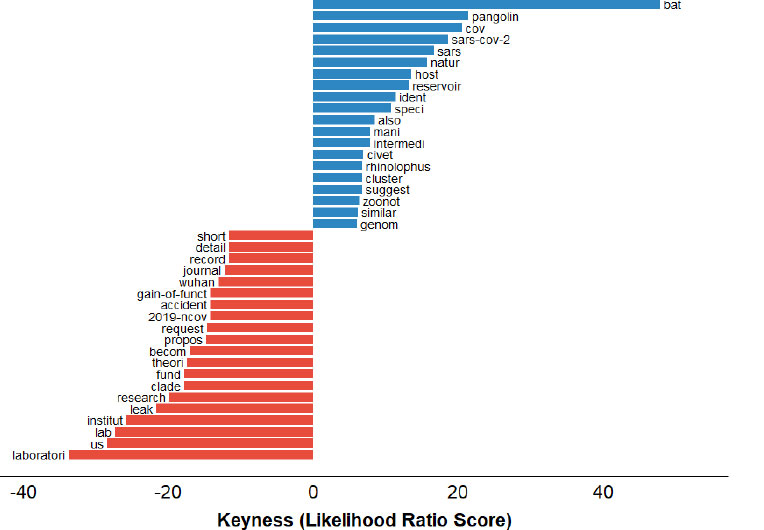
Terminology related to the origin theories, highlighting the natural origin by color blue and the lab origin by color red.
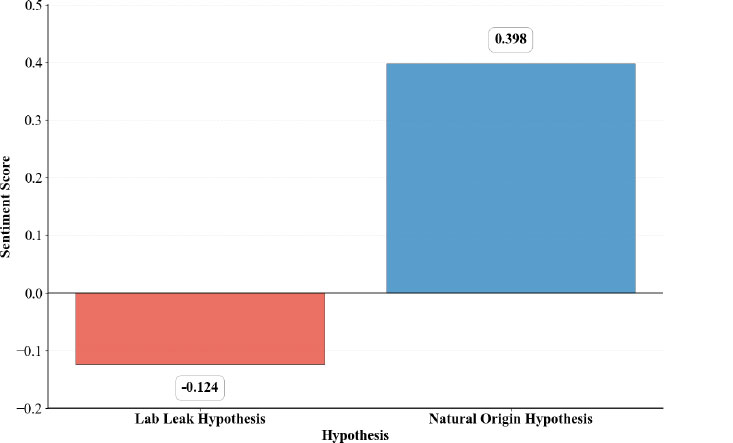
AFINN sentiment score associated with the origin hypothesis.
4.2. Scientific Evidence Supporting the Origin Hypothesis
A vast body of peer-reviewed genetic, virological, and epidemiological evidence strongly supports the hypothesis that SARS-CoV-2 emerged through natural evolutionary processes through the zoonotic spillover rather than laboratory manipulation. Several scientific statements (Segments 3, 11, 29, 263, 15, 29, 77, 178, 188, 200, 247, 155, and 357, Supplementary 11) stated that SARS-CoV-2, SARS-CoV(the virus classified as a β-coronavirus), and MERS-CoV are part of the sarbecovirus lineage, a group of coronaviruses commonly found in bats, especially species within the Rhinolophus genus (segment 186, 428, Supplementary 11) known to exist naturally and infected pangolins in Asia and Southeast Asia (Segments 2, 31, 34, 55, 56,57, 124, 228, Supplementary 11). The role of bats as reservoirs for coronaviruses, including SARS-CoV, MERS-CoV, and SARS-CoV-2, is a recurring theme across numerous research findings. Several studies demonstrated viral RNA similarities between human-infecting coronaviruses and bat viruses (Segments 68, 171, 200, Supplementary 11). Despite some hypotheses involving other animals such as snakes, genomic data have consistently ruled them out, reinforcing bats as the key reservoirs (Segment 166, Supplementary 11). It has since been established that bats host hundreds of coronavirus strains globally (Segment 6, 282, 311, 319, Supplementary 11), making them significant animal reservoirs with a broad distribution across regions like Africa, the Americas, Asia, and particularly China (Segment 8, 314, 320, Supplementary 11) which the scientists have long warned of the danger this pose to human population (Segment 1, 9, 34, Supplementary 11).
The sarbecovirus isolated from bats (Rhinolophus malayanus) in Laos was stated to exhibit high genomic similarity with SARS-CoV-2, with one particular strain, BANAL-52, showing about 96.8% similarity at the whole genome level (segment 328, Supplementary 11), while RaTG13, another bat coronavirus found in Rhinolophus affinis (horseshoe bats) in Yunnan Province, shows 96.2% similarity (segment 178, 32, 35, 46, 59, 67, 126, 139, 196, 266, 267, Supplementary 11), and that more than 780 partial coronavirus sequences have been identified in bats across 41 species infected by α-coronaviruses and 31 species by β-coronaviruses (Segment 14, Supplementary 11). Phylogenetic analyses and protein sequence alignments (segments 36, 80, 172, 207, 279, Supplementary 11) also support the close evolutionary relationship between SARS-CoV-2 and bat coronaviruses, and by the known role of bats as reservoirs for SARS-CoV and MERS-CoV (Segments 6, 8, 9, 34, 77, 162, 314, Supplementary 11). While some researchers have proposed possible intermediate hosts such as pangolins (segments 140, 265, Supplementary 11) and raccoon dogs (segment 304, Supplementary 11) based on genomic similarities (99% receptor-binding domain similarity in pangolin-CoVs) (Segments 138, 233, 446, Supplementary 11), the evidence indicates that SARS-CoV-2 likely originated from a recombination of bat-CoV-RaTG13-like viruses and pangolin-CoVs, possibly facilitated by environmental conditions that promote interspecies viral exchanges (segment 325, Supplementary 11). While definitive proof of pangolins as the intermediate host is lacking (segments 73, Supplementary 11), the presence of SARS-CoV-2-like viruses in smuggled pangolins from Southeast Asia (segments 379, 394, 448, 353, 226, 234, 538, Supplementary 11) and the possibility of cross-species transmission through wildlife markets or transport routes (segments 440, 452, 439, Supplementary 11) support this hypothesis. However, these species have shown both infection and antibody responses (Segment 138, Supplementary 11). Studies also suggest that recombination events between bat and pangolin viruses may have facilitated the emergence of SARS-CoV-2 (segments 252, 514, Supplementary 11). Although genomic studies lean more heavily toward bats as the original host (segments 505, 538, Supplementary 11), pangolins remain a significant focus due to their high genetic similarity to the virus and their documented infection with related strains (segments 583, 265, 445, 449, Supplementary 11).
Ecological disruptions and wildlife trade have been identified as significant factors in promoting cross-species transmission (Segments 12, 41, 309, 323,326, 362,364, Supplementary 11), as well as dense cave populations (Segments 324, 325, Supplementary 11), which create an ideal condition for viral evolution and spillover. Based on science findings, bats are known to be the second-highest number of mammal species after rodents. They are particularly adept at hosting zoonotic viruses due to traits like dense populations and high mobility (Segments 44, 316, Supplementary 11). Epidemiologically, the notion that SARS-CoV-2 originated through zoonotic spillover is strongly supported by a range of scientific literature and investigations (While direct bat-to-human transmission is theoretically possible (Segments 436, 437, Supplementary 11), with multiple segments emphasizing this path of transmission (Segments 134, 140, 226, 272, 381, Supplementary 11). Historical data support this, with evidence from the 2002-2003 SARS noting the bat-to-camel-to-human transmission route for MERS and the possible involvement of civet cats in the spillover of SARS-CoV (segment 19, 155, 357, Supplementary 11). It was stated that primates such as macaques could plausibly serve as intermediate hosts for SARS-CoV-2 due to their close genetic relationship with humans (segment 48, Supplementary 11). This aligns with the assertion in segment 52(Supplementary 11) that SARS-CoV-2 likely originated as an animal coronavirus that eventually adapted for human-to-human transmission. Strengthens this link, a connection between the first COVID-19 patients and the Wuhan wildlife market was observed (Segment 444, Supplementary 11), supported by positive environmental samples and genetic traces from early cases (Segments 97, 309, 370, 371, 375, Supplementary 11). This type of live animal market event had been recorded in the past (Segment 371, Supplementary 11), while the likelihood of direct transmission to scientists from non-bat wildlife species has been dismissed (Segment 472, Supplementary 11)
Importantly, SARS-CoV-2 lacks genetic fingerprints associated with laboratory manipulation (segments 23, 25, 107, Supplementary 11). The furin cleavage site in SARS-CoV-2’s spike protein, once cited as potential evidence of genetic engineering, is now understood to occur naturally via recombination, as it has also been identified in other coronaviruses (Segments 26, 140, 171, 250, 288, 547, Supplementary 11) supporting the idea that such insertions can emerge through recombination and natural selection (Segments 251, 289, Supplementary 11). Moreover, mutations such as N501Y are consistent with natural adaptation rather than artificial insertion, enhancing the virus’s transmissibility (Segment 293, Supplementary 11). SARS-CoV-2 lacks any genetic markers indicative of laboratory manipulation (Segments 23, 25, 107, Supplementary 11). While early concerns over engineered features existed, many scientists now agree that the weight of current peer-reviewed evidence points to a natural spillover event from animals to humans, likely originating in bats and possible involving intermediate hosts like pangolins or civets (Segments 78, 81, 100, 105, 144, 204, 252, 471, 473, 547, 548, 561, 104, 112, 174, 185, Supplementary 11). Although the evidence remains partly circumstantial, the majority scientific consensus supports natural emergence as the most plausible explanation for the origin of SARS-CoV-2 (Segments 116, 191, 241, Supplementary 11). The joint WHO–China report (Segments 97, 98, 115, 300, 313, Supplementary 11) deemed a natural zoonotic spillover “likely to very likely,” while a lab-related incident was labeled “extremely unlikely.” Multiple intelligence agencies concluded that a natural zoonotic spillover was the probable origin of the virus (Segment 98, Supplementary 11). Although some questions remain, the scientific consensus overwhelmingly favors a natural emergence, rooted in bat reservoirs, with possible contributions from intermediate hosts like pangolins, raccoon dogs, or civets (Segments 100, 105, 144, 204, 252, 247, 262, Supplementary 11).
4.3. Politicization of the COVID-19 Origin Debate
Early in the pandemic, the Trump administration raised the possibility that the virus (SARS-CoV-2) may have originated from a lab in Wuhan, China (Segment 5, Supplementary 12). This theory gained varying levels of credence, with the U.S. Department of Energy and the FBI later expressing “low” and “moderate” confidence, respectively, in the lab-leak hypothesis (Segment 21, Supplementary 12), while other agencies remained inconclusive, citing the need for further evidence and cooperation from China (Segment 22, Supplementary 12). Tension was raised with a reciprocal accusation. The Chinese Ministry of Foreign Affairs made an unsubstantiated claim that the U.S. military brought the virus to China, which prompted counter-accusations from then-President Trump and led to the U.S. initiating withdrawal from the World Health Organization (WHO) (Segment 6, 9, Supplementary 12). In 2021, the need to respond to the politicization arose, and President Biden ordered an intelligence review and later declassified related documents. Later, in 2023, additional documents related to the matter were declassified (Segment 19, 20; Supplementary 12). Since the issue of discovering the origin of COVID-19 became politicized in the U.S., with the Republican lawmakers holding congressional hearings targeting officials like Dr. Anthony Fauci and agencies involved in pandemic-related research (Segment 14, 15, Supplementary 12). Moreso, the U.S. agencies accused China of withholding data and destroying virus samples for investigation (Segments 23, 25, Supplementary 12). China, in turn, denied the allegation and claimed the U.S. was politicizing science (Segment 26, Supplementary 12). This resulted in a widened geopolitical conflict, causing China to impose trade sanctions on Australia after it called for an independent investigation (Segment 11, Supplementary 12), and further revelations showed that U.S. intelligence had run social media campaigns to discredit Chinese vaccines and equipment (Segment 32, Supplementary 12).
Despite the effort of the WHO trying to get all important information from China, the data retrieved was limited until after three years. Some data was finally released, but it was quickly taken down from public websites (Segment 27, 28, Supplementary 12). Experts criticized China for this delay and lack of cooperation, which increased international mistrust (Segment 24, 30, Supplementary 12). Rumors also grew regarding the Wuhan Institute of Virology (WIV), creating speculation about the illness of WIV researchers before the pandemic’s official onset, and its connection to the Chinese military (Segments 16, 17, 18, Supplementary 12). The debate continues to be shaped as much by political interests and conspiracy theories as by scientific inquiry, illustrating how the pandemic’s origins have become a battleground for geopolitical rivalry and public accountability. The scientists are very concerned about the politicization of the COVID-19 pandemic because the findings should be a matter of science, not politics (D185 and D186, Supplementary 12). Within the U.S., the government also criticized its own health agencies, like the CDC and FDA, for making poor decisions during the crisis (Segment 74 to 105, Supplementary 12).
The U.S. government report (D186, Supplementary 12) claims that Chinese officials promoted unlikely explanations for the origin of COVID-19, such as the virus spreading through frozen seafood or coming from U.S. laboratories (segment 107, Supplementary 12). A pattern of suppression and censorship was also reported by the U.S. government, claiming researchers like Professor Zhang, who first identified and sequenced the virus, were silenced and had their laboratories closed (segment 110, Supplementary 12). The report stated that there’s evidence that Chinese authorities banned the sharing of outbreak data (segments 113, Supplementary 12) and even ordered the destruction of early virus samples (Segment 114, Supplementary 12). It also mentions that whistleblowers were cracked down on (segment 115, Supplementary 12) and online discussions were censored (segments 112, Supplementary 12). Notably, the report suggests that the Chinese government focused more on political control than on being open and transparent, although no strong evidence was provided to fully support these claims.
4.4. Global Health, Policy Implications, and Impact on the Scientific Community
The impact of the politicization of the origins of COVID-19, as well as earlier outbreaks like SARS, has influenced the global health responses, public confidence, and international relationships. Although scientists and global health organizations tried their best to investigate the origin of the virus, American politicians, especially during the Trump administration, used the pandemic crisis to encourage anti-China sentiment, even voting to demand financial reparations from China and threatening to cancel U.S. debt obligations [54]. These moves, backed by significant public support, often lacked scientific grounding and ignored the complexity of zoonotic disease emergence. This politicization resulted in conspiracy theories, distracted from genuine scientific investigations, and increased international tensions. Also, scientists such as Dr. Anthony Fauci became targets of personal attacks, and private scientific communications were misrepresented to create controversy about the origins of COVID-19. Attempts were made to carry out neutral investigations, including the Biden administration’s request for intelligence agencies to examine the origins of SARS-CoV-2, which occurred in a highly politicized environment, causing misinformation to spread by social media globally [55]. As a result, the investigation into COVID-19’s origins has faced greater obstacles than the 2003 SARS outbreak, which was traced to be transferred from animals to humans through rapid and cooperative scientific efforts [56, 57]. The investigation into the origin of COVID-19 has been significantly hindered by politicized rhetoric and blame-shifting. This requires the need for scientific inquiry without political interference to better prepare and respond to future pandemics. Beliefs about where COVID-19 originated have also strongly influenced public opinion, health behaviors, and policy decisions. Unlike HIV, which required political activism to gain attention, COVID-19 became heavily politicized early on, with intelligence agencies contributing to public uncertainty. This confusion led many people to encourage the spread of conspiracy theories [58, 59, 7], resulting in the proliferation of conspiracy theories. This belief has led to reducing support for public health measures like mask-wearing and hand hygiene [7], and public attitudes and behaviors, including policy preferences; for instance, people who believe in a lab origin are more likely to support punitive measures against China [7], while those accepting a natural origin tend to advocate for increased funding for zoonotic virus research [7]. Moreover, misinformation and competitive media framing can undermine scientific consensus and long-term public trust in science [60, 61, 62]. However, the consequence of the politicization of science will not only threaten COVID-19 studies but also preparedness for future pandemics. Therefore, advancing our understanding of host–sarbecovirus co-evolution and the receptor usage determinants of these viruses becomes essential for informing evidence-based global health strategies [63]. Ultimately, the framing and communication of COVID-19’s origins have significant “downstream effects” on public policy and global health strategies [64].
RECOMMENDATIONS AND CONCLUSION
This body of evidence strongly points to a natural zoonotic origin for SARS-CoV-2. The genetic information, in combination with ecological observations and epidemiological patterns, presents a coherent picture of viral emergence through well-documented natural processes. However, history showed that diseases like HIV, Ebola, and SARS are difficult to find the precise animals or intermediate host responsible for such pathogens, and sometimes we never get a definite answer because it often takes years. Even then, definitive conclusions are not always achievable. These challenges are made even harder when scientific investigations run into political obstacles, such as China's alleged lack of transparency, which some argue may prevent us from ever achieving certainty about SARS-CoV-2’s origins. Determining the origins of COVID-19 remains complex and will require prolonged global cooperation with sustained scientific inquiry as an essential step forward.
In this study, we acknowledged certain limitations such as gaps from inaccessible data, a potential language bias from relying on English-language potentially omitting important regional or non-Western perspectives, and the evolving nature of the evidence itself. Some government reports and origin-related research may remain classified or inaccessible, introducing gaps. Due to the evolving nature of COVID-19 origin debates, it’s possible that new evidence may alter the current conclusions. Despite these limitations, our mixed-method meta-analysis offers a robust and nuanced framework for examining the COVID-19 origin debate in its full scientific and social context [Supplementary 2]. Therefore, it is essential for further studies to apply empirical data to examine how the politicization of the COVID-19 origin debate has influenced scientific communication, research integrity, and public trust. These areas remain underexplored and warrant systematic investigation to better understand the broader implications of contested scientific narratives.
AUTHORS’ CONTRIBUTIONS
The authors confirm their contribution to the paper as follows: A.C.O., O.S.O., O.C.K.: Study conception and design; B.K.O, S.K.O., A.A.T., A.A.A.: Data collection; A.J.D., O.M.I.: Data Analysis or Interpretation; F.O.O., E.B.N.: Validation; A.E.J., R.C.O., S.H.S., I.G.E., A.S.A., A.Y.A., A.T.I., S.O.O.: Draft manuscript. All authors reviewed the results and approved the final version of the manuscript.
LIST OF ABBREVIATIONS
| WIV | = Wuhan Institute of Virology |
| WHO | = World Health Organization |
ACKNOWLEDGEMENTS
Declared none.
SUPPLEMENTARY MATERIAL
Supplementary material is available on the publisher’s website along with the published article.
Supplementary files
Supplementary 1: Synthesized documents metadata.
Supplementary 2: PRISMA 2020 Checklist document.
Supplementary 3: Source Documents for Thematic Analysis (Anonymized as D1, D2, ... Dn).
Supplementary 4: Study-Level and Author-Level Metadata Sheets (Excel Format).
Supplementary 5: Definitions of Thematic Categories Used in the COVID-19 Origin Analysis.
Supplementary 6: Jaccard Similarity Scores.
Supplementary 7: Levenshtein Distance Scores.
Supplementary 8: Supplementary analysis results including Tables S1-S4 and figures Figure S1-S3
Supplementary 9: TF-IDF Weighted Keyword Tables for Competing Hypotheses.
Supplementary 10: Keyness Analysis Output: Term Salience in Competing Hypotheses.
Supplementary 11: Thematic Segments Citing Scientific Support for Origin Hypotheses.
Supplementary 12: Thematic Segments Highlighting Political Influences on the Origin Debate.
