All published articles of this journal are available on ScienceDirect.
SARS-CoV-2 Therapeutic Landscape, Opportunity and Future Threats
Abstract
During the past two decades, the world has seen several known and novel zoonotic viruses and deadly bacterial diseases, such as West Nile Virus (1999 to 2002), Anthrax (2001), H1N1(2009), Ebola (2014), Zika Virus (2016), SARS-CoV (2002), MERS-CoV (2012) and SARS-CoV-2 in 2019. The current ongoing COVID-19 pandemic is completely unpredicted and it has hugely changed our health care systems, global economy and social lifestyles. SARS-CoV-2 is still under genetic evolution and getting mutated to escape our immune system and showing resistance against available therapies. In this current research work, we have examined all publicly available scientific literature to date to understand the genetic evaluation of coronavirus species and their transmission possibilities to humans. We have also explored recently reported mutations of concerns in viral spike glycoprotein. We then discussed various SARS-CoV-2 preclinical and clinical research breakthroughs and highlighted our limitations and readiness to combat this deadly disease. Based on our recent study, we have emphasized developing a global viral, fungi and microbes platform. It can help us to predict mutations on their genomic, structural and pathophysiological profile to better address early on future threats by such infectious agents.
1. INTRODUCTION
The “severe acute respiratory syndrome coronavirus 2” (SARS-CoV-2) also known as COVID-19 is a highly infectious virus able to cause a novel form of viral pneumonia that has become the number one health challenge around the world [1]. It was first reported in late 2019 in Wuhan, China and has since spread extensively worldwide and was named by WHO on 12 January 2012 [1, 2]. Considering the severity of symptoms among the affected people and rapid spread, the World Health Organization (WHO) declared COVID-19 as a pandemic on March 11th, 2020. SARS-CoV-2 is a single-stranded enveloped ribonucleic acid virus with an average size of 75 to 150 nm and it belongs to the subfamily Orthocoronavirinae in the family Coronaviridae, Order Nidovirales [3]. It has shown almost 85% structure similarity with SARS-CoV and the main functions of their structural proteins, such as spike (S) surface glycoprotein, membrane (M) protein, envelope (E) protein, and nucleocapsid (N) protein, as well as nonstructural proteins, such as RNA polymerase (RdRp), papain-like protease (PLpro) coronavirus main protease (3CLpro) are also highly conserved [4]. Phylogenetic analysis shows bat coronavirus RaTG13 is the most similar and closely associated virus, a virus sampled from a horseshoe bat (Rhinolophus affinis) belonging to this suborder [5].
As of today 28th June, 2021, 179,686,071 confirmed cases of COVID-19, including 3,899,172 deaths, were reported by WHO (https://covid19.who.int/). Till today, 22.6% people have received at least one dose of a COVID-19 vaccine (https://ourworldindata.org/covid-vaccinations). Although most of the available vaccines are capable of preventing COVID-19 infections to a high percentage, still COVID-19 positive cases are observed in the individuals who had received their either first or second dose of vaccines [6, 7]. Scientists at the Indian Council of Medical Research (ICMR) reported COVID-19 cases are 0.04% in India after vaccination. A genome sequence comparison study conducted by the Institute of Genomics and Integrative Biology (IGIB) revealed that in most of these cases COVID-19 delta variant (B.1.1.7) was common with E484K and S477N mutations in spike protein, which help these variants to escape from our immune system [8].
In total, 122 vaccines are in the preclinical development stage and 92 vaccine candidates are being evaluated in clinical trials (https://biorender.com/covid-vaccine-tracker). BNT162b2 (Pfizer/BioNTech) and AZD1222 (Oxford-AstraZeneca) were the first companies to file for an emergency use authorization with the United States Food and Drug Administration (FDA) for their coronavirus vaccine candidate. Meanwhile, mRNA-1273 (Moderna, Inc), Ad26.COV2.S (Johnson and Johnson), CoronaVac (Sinovac Biotech) and BBIBP-CorV (Sinopharm) also got their COVID-19 vaccines approved by FDA. In addition to SARS-CoV-2 specific vaccines, investigations of Measles-Mumps-Rubella vaccine [9, 10], Oral Polio Vaccine [11] and Bacille Calmette-Guerin vaccines for their repurposing potential against COVID-19 [12] are being conducted under NCT04333732, NCT04540185 and NCT04379336 clinical trial, respectively.
We leveraged Ontosight® Discover, an artificial intelligence based semantic discovery platform which understands biomedical concepts and allows us to identify relevant results in context across multiple data sources like publications, clinical trials, congresses, theses, patents, grants, and publicly available databases (https://ontosight.ai/). We did explore all PubMed indexed journals through this platform to analyze the therapeutic landscape and limitations of coronavirus research. In this current review article, we have explored all reported variants of SARS-CoV-2, their origin and explored various in-silico technologies, which are used to accelerate drug development. This review also alerts and recommends research areas with the need to prepare for future viral and biological threats.
2. CORONAVIRUS IS NOT A NEW VIRUS
Numerous coronaviruses were discovered and reported in poultry around the 1930s, causing respiratory, gastrointestinal, liver, and neurologic diseases in animals. The sub-family of Coronaviridae, which contains viruses of both medical and veterinary importance, can be divided into the four genera alpha (α), beta (β), gamma (γ) and delta coronavirus (δ) that were at approximately 2400 base count (BC), 3300 BC, 2800 BC, and 3000 BC, respectively [13].
Seven coronaviruses (α and β) known to cause symptomatic diseases in humans are mentioned in Table 1. Four coronaviruses (Coronaviruses 229E, OC43, NL63, and HKU1) cause about 15% to 30% of cases of the common cold and pneumonia in human beings. The remaining 3 coronaviruses (SARS-CoV-2, MERS-CoV, SARS-CoV) cause much more severe, and sometimes fatal, respiratory infections in humans than other coronaviruses and have caused major outbreaks of deadly pneumonia in the 21st century.
| Serial Number | Species | Coronavirus (Genus) | Diseases | Taxonomy ID | |
|---|---|---|---|---|---|
| Alphacoronavirus (α) | |||||
| 1 | Human | HCoV-229E | Common cold | 11137 | |
| 2 | Human | HCoV-NL63 | pneumonia and bronchiolitis | 277944 | |
| 3 | Pig | TGEV | Gastroenteritis | 11149 | |
| 4 | Pig | Porcine respiratory coronavirus (PRCoV) | Mild respiratory disease | 11146 | |
| 5 | Pig | Porcine epidemic diarrhea (PEDV) |
Gastroenteritis | 28295 | |
| 6 | Cat | Feline infectious peritonitis (FIPV) |
Peritonitis, pyogranuloma | 11135 | |
| 7 | Cat | Feline coronavirus (FECV) | Subclinical enteric disease | 12663 | |
| 8 | Dog | CCoV | Enteritis | 11153 | |
| 9 | Bat | Rhinolophus bat coronavirus HKU2 (Rh-BatCoV-HKU2) | - | 693998 | |
| 10 | Bat | Miniopterus bat coronavirus HKU8 (Mi-BatCoV-HKU8) | - | 694001 | |
| 11 | Bat | Mi-BatCoV 1B | - | 694000 | |
| 12 | Bat | Sc-BatCoV-512 | Scotophilus bat coronavirus | 693999 | |
| Betacoronavirus (β) | |||||
| 13 | Human | SARS-CoV-2 | COVID-19 | 2697049 | |
| 14 | Human | SARS-CoV | Severe acute respiratory syndrome | 1431340 | |
| 15 | Human | MERS-CoV | Middle East respiratory syndrome | 1335626 | |
| 16 | Human | HCoV-OC43 | Common cold | 31631 | |
| 17 | Human | HCoV-HKU1 | Common cold | 290028 | |
| 18 | Pig | PHEV | Vomiting and wasting disease, encephalomyelitis | 42005 | |
| 19 | Pig | SADS-CoV | Gastroenteritis, diarrhea | 2032731 | |
| 20 | Rat | Rat sialodacryoadenitis coronavirus (SDAV) | Sialodacryoadenitis, mild pneumonia | 92931 | |
| 21 | Rat | Rat coronavirus Parker (RCoV-P) | Fatal pneumonia | 502102 | |
| 22 | Horse | ECoV | Mild gastrointestinal disease | 136187 | |
| 23 | Bovine | BCoV (Bovine coronavirus) | Enteritis, respiratory disease, pneumonia | 11128 | |
| 24 | Bat | Pi-BatCoV-HKU5 | - | 694008 | |
| 25 | Bat | Ty-BatCoV-HKU4 | - | 694007 | |
| 26 | Bat | Ro-Bat-CoV HKU9 | 694006 | ||
| 27 | Bat | SARSr-Rh-BatCoV HKU3 | Severe acute respiratory syndrome | 333387 | |
| 28 | Human | Civet SARS CoV SZ16/2003 (SARSr-CiCoV) | Severe acute respiratory syndrome | 231515 | |
| 29 | Human | Civet SARS CoV SZ3/2003 | Severe acute respiratory syndrome | 231513 | |
| 30 | Equine | Equine Coronavirus (ECoV) | - | 136187 | |
| 31 | Giraffe | GiCoV | Severe diarrhea | ||
| 32 | Rat | RCoV | Severe acute respiratory | 31632 | |
| 33 | Antelope | Sable antelope coronavirus US/OH1/2003 (AntelopeCoV) | - | 422138 | |
| 34 | Mouse | MHV | Murine coronavirus MHV-1 | 502106 | |
| 35 | Dog | Canine respiratory coronavirus (CRCoV) | Mild to severe respiratory disease, pneumonia | 215681 | |
| Gammacoronavirus ((γ)) | |||||
| 36 | Chicken | IBV | Bronchitis, nephritis | - | |
| 37 | Turkey | TCoV | Contagious enteric disease in turkeys | 19184806 | |
| 38 | Whale | BWCoVSW1 | Pulmonary disease, terminal acute liver failure | - | |
| Deltacoronavirus (δ) | |||||
| 39 | Pig | Porcine deltacoronavirus (PDCoV) | Gastroenteritis | 1586324 | |
| 40 | Bulbul | HKU11 | Respiratory disease | 574549 | |
| 41 | Avian | HKU12 | Thrush coronavirus | 1297662 | |
| 42 | Avian | HKU13 | munia coronavirus | 1297661 | |
| 43 | Porcine | PorCoV HKU15 | undifferentiated febrile illness | 1159905 | |
| 44 | Vertebrates | WECoV HKU16 | White eye coronavirus | 1159907 | |
| 45 | Sparrow | sparrow coronavirus HKU17 (SpCoV HKU17) | Respiratory disease | 1159906 | |
| 46 | Avian | MRCoV HKU18 | Magpie robin | 1159903 | |
| 47 | Night heron | NHCoV HKU19 | - | 1159904 | |
| 48 | Wigeon | Wigeon coronavirus HKU20 (WiCoV HKU20) | - | 1159908 | |
| 49 | Avian | CMCoV HKU21 | Common moorhen | 1159902 | |
The human Alphacoronavirus (HCoV-229E and HCoV-NL63) and Beta-coronavirus (SARS-CoV-2, SARS-CoV, MERS-CoV, HCoV-OC43 and HCoV-HKU1) are derived from bat and have a high genome similarity with bat viruses [14] whereas, Gamma-coronavirus (γ) and Delta-coronavirus (δ) are mostly derived from birds and pigs [15]. Recent reports suggest a transmission to humans as well (specific antibodies were found in human plasma samples in Haiti) [16]. Crosslinking and transmission of these viruses across mammal species is an evolutionary process [17].
Recent alteration of human microbiota in SARS-CoV-2 patients and secondary fungal and bacterial infections during the treatment of COVID-19 were described [18]. Since viral infection increases the susceptibility for secondary infections, it is also important to develop a strategy and guideline to deal with classes of viruses and other micro-organism together. Interactions and complications due to viruses, bacteria and fungi together can lead to critical clinical conditions in the future [19].
3. FREQUENT MUTATIONS AND FUTURE THREATS
Multiple SARS-CoV-2 lineages and spike protein mutations were reported across the globe, such as Epsilon [20, 21], Zeta [22], Eta [23], Theta [24, 25] (https://pgc.up.edu.ph/covid19-p3-theta-who/), Iota [26] and Kappa [27]. These variants mainly affect the receptor binding domain (RBD). WHO has classified these mutations into two groups; the first group is “Variants of Interest”, which includes Eta (B.1.525), Theta (P.3), Iota (B.1.526, B.1.526 and B.1.617), Zeta (P.2) and Kappa (B.1.617.1 and B.1.617.3) lineage and “Variants of Concern”, which includes Alpha (B.1.1.7), Beta (B.1.351), Epsilon (B.1.427, B.1.429), Delta (B.1.617.2) and Gamma (P.1).
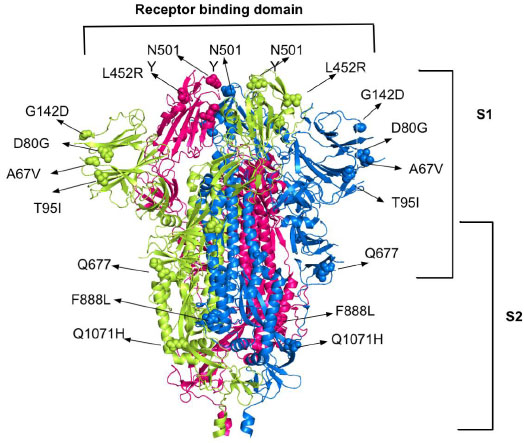
The most common mutations in spike proteins for the ACE2 receptor are A67V, N501Y, K417, L452R and Q677, F888L, T95I, D80G, D950H, Q1071H and G142D. Deleterious mutations in coronavirus spike protein, such as D614G, E484K, P681R, E154K, P681R, and V1176F were also found. These mutations on spike proteins help them to escape the human immune system and bind to the human host cells which leads to the resistance of available therapy and combination therapy especially with monoclonal antibodies [27, 28]. A recent study shows more than 44 different mutation locations in spike proteins [29]. The B.1.525, B.1.526, B.1.526.1, B.1.617, B.1.617.1, B.1.617.3, P.2 mutations are all over the proteins, including receptor binding domain (RBD), which makes its inhibition more challenging (Fig. 1).
4. MAJOR BREAK THROUGH OF COVID-19 RESEARCH
4.1. Genome Sequencing and Characterization
During the initial weeks of COVID-19 spread, the world had major challenges to diagnose, characterize and understand SARS-CoV-2’ virulence and pathogenesis mechanism. The first major breakthrough was achieved after the successful characterization of SARS-CoV-2 [30]. A research team of Fudan University published the complete genome of SARS-CoV-2 publicly online so that it could be utilized further for quick research and further prevention strategies and therapeutic development [31]. The sequence reads generated from this study are available at GenBank under accession number MN908947.
4.2. Tertiary Structure of SARS-CoV-2 Proteins
We searched for SARS-CoV-2 proteins in the Protein data bank (PDB) using a search keyword “COVID-19” in Source Organism Taxonomy Name (Full Lineage) on 28th June 2021. It contains 1,277 COVID-19 proteins that have the taxonomy name as “SARS-CoV-2”. In total, 284 human coronavirus protein structures were identified in PDB, which include X-ray diffraction (122) and electron microscopy (172). The first protein structure of SARS-CoV-2 main protease (PDB ID:6LU7) that was expressed in Escherichia coli BL21(DE3) was deposited on 26th January, 2020 and annotated and published in PDB for immediate further research [32]. On 06 February, 2020, a watercolor of Coronavirus was published by David S. Goodsell and the team described coronavirus entry mechanisms in the lungs [33]. These experimental structures put a base on the early drug discovery on COVID-19.
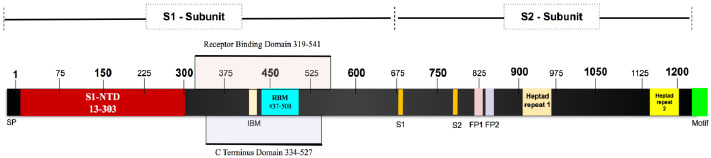
The SARS-CoV-2 Spike glycoprotein is a 1281 amino acid protein. It is a homotrimer and each monomer consists of two domains, S1 and S2 to make one complete spike protein [34]. The extracellular N-terminal domain S1 (14-685) helps in recognition and binding to the host ACE receptor and internalization of the virus into the endosomes of the host cells [35, 36]. Intracellular C-terminal S2 (686-1273) during virus endocytosis S2 acts as a class I viral fusion protein and mediates fusion of the complete viral genetic materials into the host cellular membranes [37]. It contains two heptad repeats, which is an ideal feature of the virus to function as a fusion protein (Fig. 2).
5. VARIOUS EXPERIMENTAL AND COMPUTATIONAL APPROACHES IN SARS-COV-2
We queried Ontosight Discover® using our life science ontology for SARS-CoV-2, which includes synonyms of SARS-CoV-2 as “2019ncov, disease caused by 2019-ncov, coronavirus disease2019, sars-cov-ii disease, COVID-19, 2019-ncov infection, sars-cov-2 disease, human coronavirus 2019 (hcov-19), hcov19, severe acute respiratory syndrome coronavirus 2, corona virus disease (COVID-19), novel corona virus 2019, 2019 ncov, wuhan coronavirus infection, wuhan seafood market pneumonia virus infection, 2019ncov infection and novel corona virus2019” and searched for only PubMed Indexed articles from December 1st, 2019 to the date of study i.e, 28th June 2021. It results in 149,167 articles that had SARS-CoV-2 related concepts and any one of the synonyms present in their title, abstracts, keywords or in the full-text body. Out of these, 18,805 articles were under the “Review” category, 113,888 articles were under “Journal articles” and 7,690 articles were published as a case report while the remaining 8,784 belonged to the other categories such as congress, meta-analysis, etc. We explored these articles to understand the contribution of various wet-labs (laboratories where biological molecules or samples are being analyzed using liquids) and dry lab (laboratory where computational or applied mathematical analyses are done with the assistance of computer-generated models) techniques associated with COVID-19 to understand the SARS-CoV-2 research gaps and identify future viral threats.
Our study revealed that experimental studies (wet lab) were published more in number when compared to computer-based approaches. Most of the experiments were conducted to understand the virus pathogenesis and virus itself, such as molecular assays for antiviral activity, microscopy and imaging techniques and biochemical analysis, whereas experimental protein structure determination was conducted to understand viral proteins. Moreover, expression and multi-omics analysis were reported and studied in various publications for further therapeutic development (Fig. 3).
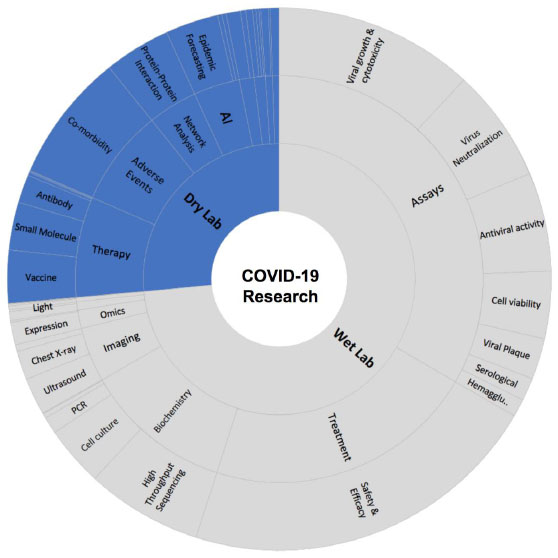
Computational biology and the role of artificial intelligence also played a crucial role to uncover various pathophysiological processes and the origin of this virus. Our today’s understanding about COVID-19 and recent therapeutic developments is an excellent example of collaborative research. Due to the advance development of artificial intelligence, such as neural networks, machine learning, chemoinformatics, system biology, network analysis and large-scale genomics analysis, it was possible to identify more than 141 repurposable drugs to be tested against COVID-19 (Table S1).
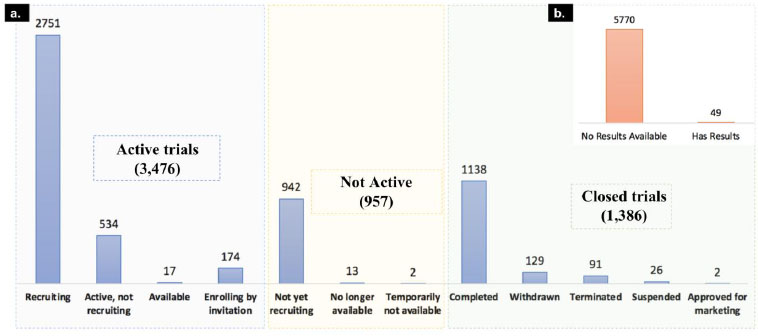
As of today, 5,819 clinical trials are registered in clinicaltrials.gov for COVID-19 and associated disease. For 5,770 of these clinical trials, the results are yet to be published. We classified the clinical trials into 3 major classes based on their recruiting status. The first category is active trials; it includes clinical trial status with “recruiting”, “active, not recruiting”, “enrolling by invitation” and “available” trial phases. The second category is not active, which includes “not yet recruiting”, “no longer available”, and “temporarily not available” clinical trials and the third category is Closed trials, which includes “completed”, “withdrawn”, “terminated”, “approved for marketing” and “suspended” as mentioned in Fig. (4).
The majority of the trials are in the active stage, which suggests a number of drug candidates are under investigation to evaluate their treatment benefits against SARS-CoV-2. We downloaded all these clinical trials and studied investigated drugs and their mechanism of actions to understand the current therapeutic landscape of SARS-CoV-2.
We identified 373 unique therapeutic agents that are under clinical trials, out of which 141 drugs are repurposable drugs (already approved drugs for other disease), while remaining 232 drugs are either novel therapeutics (specifically proposed for COVID-19) or not approved for any indication yet. In total, 107 drugs reached clinical phase 4 (approved for marketing) against various clinical studies associated with COVID-19 (Table S2). Until now, only one drug (Remdesivir) is approved against COVID-19, while for the 3 other drugs, an emergency use authorization has been granted for mild to moderate SARS-CoV-2 infections, as shown in Table 2.
| Drug Name | MoA | Approval Date | Treatment |
|---|---|---|---|
| Remdesivir | RNA-dependent RNA polymerase (RdRp) | October 22, 2020 | Mild, moderate Novel Coronavirus Infectious |
| Bamlanivimab & Etesevimab (LY3819253) |
Spike glycoprotein | February 9, 2021 | Mild, moderate Novel Coronavirus Infectious |
| Casirivimab & Imdevimab (Regen-COV) |
Receptor binding domain of the spike protein of SARS-CoV-2 | November 21, 2020 | Mild, moderate Novel Coronavirus Infectious |
| Sotrovimab | Virus internalisation inhibitors | May 26, 2021 | Mild-to-moderate COVID-19 in adults and pediatric patients; not shown benefit in patients hospitalized due to COVID-19 |
We identified 373 therapies (303 candidate drugs, 15 combinations, 55 other therapies) from 5,819 COVID-19 related trials (Table S3). In order to understand the effort and focus of the pharmaceutical industry for therapeutic development against various stages of COVID-19, we classified these drugs into 3 major classes based on their therapeutic use in SARS-CoV-2.
(1) Early infection phase is also known as the incubation period as in this phase, SARS-CoV-2 multiplies and develops resistance in the host. Infection starts when an uninfected person inhales COVID-19 containing droplets or aerosols and the virus interacts with angiotensin-converting enzyme 2 receptors on human cells [38].
(2) Pulmonary phase: The pneumonia phase is divided into two, depending on the absence or presence of hypoxia; whereby hypoxic pneumonia patients will likely require hospitalization and oxygen supplementation.
(3) Hyperinflammatory phase is the phase where patients struggle to take a breath. It is the most severe stage of this viral infection. An increase in several biomarkers, such as IL2, IL6, IL7, TNF-alpha and C-reactive protein, indicate this stage [39]. Anti-inflammatory drugs, immunomodulators including corticosteroids and interleukin/cytokine inhibitors are recommended at this stage. This is the most critical stage with the highest lethality among the three phases.
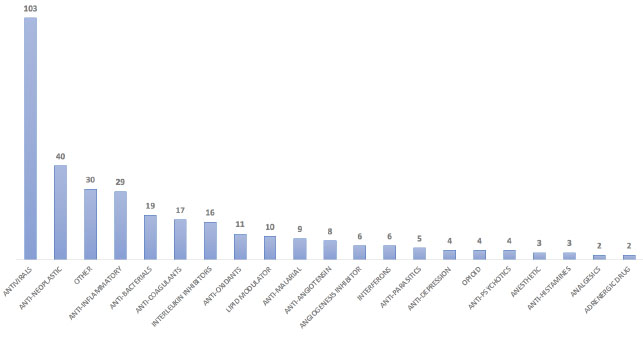
Further, 22 different mechanisms of actions were prepared based on their treatment type (Table S3). Our analysis revealed that most of the ongoing/failed clinical trials are for the early infection phase of the SARS-CoV-2 (136 drugs) followed by the hyper-inflammatory phase (119) to improve the standard of care and reduce morbidity and mortality. Eighty-one drugs are used in stage 2 to improve the medical condition of infected patients (Table 3). We identified 49 monoclonal antibodies out of these antibodies are mainly developed as virus internalization agents (Table S4) to be used as passive immunological agents to minimize virulence in mild to moderate COVID-19 patients (https://www.antibody society.org/COVID-19-biologics-tracker/) (Fig. 5).
Several antiviral drugs with distinct mechanism of action, such as virus internalisation (e.g. BRII-196, and BRII-198), RNA replicase inhibitors (e.g. Galidesivir), protease inhibitors (e.g. Lopinavir and Ritonavir), virus replication inhibitors (e.g. Sofosbuvir and Voclosporin), serine protease inhibitors (e.g. Camostat mesylate and Tranexamic acid) and dipeptidase inhibitors (e.g. Brensocatib) are the major drug classes which were investigated against COVID-19 in various clinical trials. Similarly, several immunomodulators, such as complement C3 inhibitors (e.g. AMY-101 and APL-9), glucocorticoid receptor agonists (e.g. Apremilast and Aprotinin), CD20 antigen inhibitors (e.g. Ofatumumab) and T lymphocyte stimulants were also investigated against COVID-19.
Analysis of clinical studies also suggested that 40 anti-cancer drugs are also under investigation and majority of these drugs belong to MAP-kinase-activated kinase 2 inhibitors, Tyrosine-protein kinase SYK inhibitor, JAK Inhibitor and Interleukin 1 receptor antagonists, casein kinase II inhibitors.
In order to control inflammation and bacterial superinfection during the COVID-19 treatment, several antibacterial drugs, such as beta lactamase inhibitors, cell wall inhibitors, protein 50S/70S ribosomal subunit inhibitors and cell replacements drugs were also investigated in various clinical trials against SARS-CoV-2. Moreover, in order to prevent mortality due to cardiac attack due to blood clots, several anticoagulants, such as Factor Xa inhibitors, blood coagulation factor inhibitors, plasminogen activators, thrombin inhibitors are also being tested against COVID-19.
| Stage of COVID-19 | Broad Category of Mechanism of Actions | Drug Example | Mechanism in COVID-19 | Drug Count |
|---|---|---|---|---|
|
Stage 1 (Early infection phase) |
Antivirals | Remdesivir; Favipiravir; Lopinavir; Umifenovir; ADM03820; ADG20; AZD7442; Apabetalone (RVX-208) etc. |
RNA-dependent RNA polymerase (RdRp), Viral membrane fusion; Endocytosis, non-competitive binding to viral protein, etc [40, 41]. | 103 |
| Lipid Modulator | Statins; Cyclodextrins, Opaganib etc. | Controlling virus replication. Lipids can act as direct receptors or entry cofactors for all types of viruses at the cell surface or the endosomes etc [42]. | 10 | |
| Angiotensin receptor blockers | Losartan; Spironolactone etc. | Cellular entry by decreasing ACE2 receptors expression and cluster of differentiation 147 (CD147) etc [43]. | 8 | |
| Interferons | IFN-α and β | Hinder viral replication, including viral entry | 6 | |
| Antimalarial | Hydroxychloroquine, Chloroquine etc. | Anti-inflammatory and immunomodulatory activities in treating COVID-19 [44]. | 9 | |
|
Stage 2 (Pulmonary phase) |
Anti-bacterial, incl. Antibiotics | Azithromycin; Clarithromycin etc. | Secondary infection reduces the inflammatory process and modulate the immune system etc [45]. | 19 |
| Anti-parasitic | Ivermectin; Niclosamide etc. | Control viral replication etc [46]. | 4 | |
| Antioxidants | Piperine; Selenium etc. | Reduces the production of IL-1β, IL-6, TNF-α, COX-2, nitric oxide synthase-2, and NF-κB. Neutralizes free radicals, ROS, and hydroxyl radicals [47]. | 11 | |
| Anti-neoplastic (Anticancer) |
Abemaciclib; Ruxolitinib; Duvelisib; Saracatinib; Nivolumab, kinase inhibitors etc. | Inhibition of host intracellular-viral events in viral life cycle etc [48]. | 39 | |
| Anti-depression | Fluvoxamine | Reduces the production of inflammatory molecule; controlling neuropsychiatric symptoms of COVID-19 patients etc [49]. | 4 | |
| Antipsychotics | Quetiapine; Cerebrolysin etc. | Antipsychotics modulate the expression of inflammatory cytokines and inducible inflammatory enzymes [50]. | 4 | |
| Stage 3 (Hyperinflammatory) | Anticoagulants | Heparin | Anti-thrombin prevents blood clot formation during cytokine storm etc [51]. | 17 |
| Adrenergic | Berberine, Prazosin etc. | For preventing or reducing acute respiratory distress syndrome (ARDS) and other complications [43]. | 2 | |
| Anti-inflammatory | Acebilustat; Apremilast | Limiting the expression of hyper-inflammatory genes during hyperinflammatory phase [52]. | 29 | |
| Immunomodulators (Corticosteroids) | Tocilizumab | Interrupts cellular signals transduction pathway, and subsequently decreases inflammatory responses etc [53]. | 44 | |
| Interleukin Inhibitors | Clazakizumab; Levilimab etc. | To prevent severe damage to lung tissue caused by cytokine release [54]. | 15 | |
| Analgesics | Ibuprofen | Pain management during COVID-19 infection [55, 56]. | 2 | |
| Antihistamine | Desidustat; Famotidine etc. | Prevent progression to severe disease in elderly patients [57] | 3 | |
| Anesthetic | Sevoflurane; Celecoxib etc. | To control inflammation and prevent complications during cytokine storm [58] | 3 | |
| Opioid | Remimazo etc. | Ordinary functioning of the immune system and not recommended against COVID-19 [59] | 4 |
CONCLUSION
Academic and pharma industries are working together at different levels to identify the most suitable therapeutic solutions for SARS-CoV-2. As of now, academic institutions have contributed significantly to identifying genomic level information and defining various pathophysiological processes. Crosslinking and ongoing frequent mutation in COVID-19 proteins is one of the major challenge for identifying the most effective therapeutic and effective preventive strategies. However, published biochemical studies serve a very strong background to pharma industries and publicly funded institutions for further therapeutic and preventive care solutions development.
We also understand that it is an ongoing evolution process of various viruses and microorganisms that requires continuous long-term vigilance and surveillance studies to identify and address any future pandemic risks. At the same time, we need to be prepared for any further crosslinking (intermediate host and interspecies transmission) of these viruses, their future mutations and possible transmission in humans. Virus bacterial interaction could also be a major challenge in future. These interactions can also indirectly affect the host response to viral infection and make it the worst health condition [60].
At the same time, therapies against SARS-CoV-2 and pre-existing secondary infections with viruses (Metapneumovirus, Coronavirus HKUetc.), bacteria (Klebsiella pneumoniae, Acinetobacter baumannii, Streptococcus spp.) and fungi (Candida krusei, Aspergillus spp., Candida glabrata etc.) need to monitor carefully. Other complications, such as thrombosis during treatment with diabetic, malignancy, other inflammatory and cardiovascular disorders need to be improved to control the mortality rate in hospitalised coronavirus infected patients. Treatment strategies for various groups of patients need to be established, the mortality rate due to co-infections and comorbidity in hospitalised patients needs to be improved. Therefore, we recommend a personalised medicines based therapy (biomarker based patient groups selection for clinical decision making), broad spectrum preventive care solution and immune boosters to immediately address the current standard of care against this disease.
AUTHORS’ CONTRIBUTIONS
OPS, contributed to study design, data collection, data analysis, interpretation and writing and critical review. WS and JS contributed to data interpretation, writing and critical review and finalization of the paper.
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
We thank Dr. Gunjan Bhardwaj, CEO and Founder of Innoplexus AG to support and motivate us for writing this paper. I would also like to express my gratitude to Mr. Abhijit Keskar, Senior VP of technology, Innoplexus AG, Mr. Holger Hoffman, C.F.O of Innoplexus Consulting Pvt. Ltd., Mr. Amit Ananpara, M.D India, Innoplexus Consulting Pvt. Ltd. for continuous support, motivation and infrastructure facilities. Authors of this article are also grateful to the technical staff of Innoplexus Consulting Service Pvt. Ltd. Mr. Suyash Masugade, Mr. Samadhan Khatal, Mr. Sanket Shete, Mr. Rupesh Bhosale for timely technical support. We would also like to thank Mr Milind Bhore, Mr Prakash Dhandole, Miss Jyoti Mahadik and Ms Sonal Darokar for various admin support and suggestions during this period.
SUPPLEMENTARY MATERIAL
Supplementary material is available on the publishers website along with the published article.
