All published articles of this journal are available on ScienceDirect.
Genomic Variation and Treatment Strategies of COVID-19: A Descriptive Review
Abstract
Coronavirus disease 2019 (COVID-19) was spread across China and affected more than 180 countries worldwide to date. SARS-CoV-2 is a beta coronavirus that shows genomic similarity with bat coronaviruses. The intermediate source in human viral transmission is caused by dromedary camels for MERS-CoV and civet cats for SARS-CoV. Transmission of the virus from human-to-human is achieved through close contact with infected persons. The genome of the coronavirus consists of four structural proteins, including Spike (S), Membrane (M), Envelop (E), and Nucleocapsid (N) proteins. These structural proteins are encoded within the genome 3' end. The spike protein is responsible for virus attachment to the host cell surface receptors (angiotensin converting enzyme-2 receptor), resulting in fusion and subsequently cell damage. The N protein, after binding, causes RNA genomic changes. The accessory proteins present in SARS-CoV open read frames (ORFs) are very similar to COVID 19. The COVID-19 infection triggered a number of deaths and even now affecting a number of confirmed cases. Coronavirus patients are characterized by pneumonia, cytokine storms, weakened lymphocytes, lymphocytopenia, and respiratory failure. However, the lack of antiviral vaccines permits emergency clinical trials since January 2020. Recently, several anti-viral drugs are being repositioned and restructured as part of an immediate investigation. In this review, we discussed the genomic sequence of SARS-CoV-2, its different features and current therapeutic strategies to combat this serious condition.
1. INTRODUCTION
Coronaviruses are in the family of coronaviridae. Coronavirus is a positive-sense single-stranded RNA (+ssRNA) with a much smaller size (65-125 nm in diameter). Coronaviruses are primarily subdivided into four classes, such as alpha, beta, gamma, and delta. Alpha and beta mainly infect people, while birds and mammals are infected with gamma and delta. Epidemics such as SARS-CoV in 2002-2003, Middle East Respiratory Syndrome of Coronavirus (MERS-CoV) in 2012, Acute Lung Injuries (ALI), and Acute Respiratory Distress Syndrome (ARDS) in 2012 have occurred in the last two decades [1]. China informed World Health Organization (WHO) in December 2019 about one of the unfamiliar diseases that took over 1800 lives in the first 50 days. The SARS-CoV-2 situation was confirmed by the International Committee on Virus Taxonomy (ICTV) [2]. SARS-CoV (2002-2003) infected 8422 individuals with 916 deaths, with 11% mortality rate [3]. On the other side, COVID-19 infects individuals with a seven percent mortality rate [4]. The analysis clearly shows that the SARS-CoV-2 transmission rate is higher than SARS-CoV and could be due to the spike(S) genetic mutation in the receptor domain of SARS-CoV-2. Chinese people had SARS and coronavirus confirmed in 2003 [5]. In patients with acute severe respiratory syndrome, pneumonia has occurred, and bone marrow cells can be infected, which leads to thrombo- cytopenia [6]. In 2012, the Saudi Arabia population was diagnosed with a group of coronaviruses known as Middle East Respiratory Syndrome (MERS-CoV). Patients with MERS-CoV have experienced breathing disorders and kidney failure, as well. The recent SARS-CoV-2 outbreak has mild (80%) to moderate (20%) symptoms associated. The virus was identified as a novel coronavirus following a sequence-based analysis [7].
Details of viral pneumonia in infected patients were repor- ted by the Chinese National Health Commission in January 2020 [8]. The virus transmission is caused by close contact with infected individuals, exposure to respiratory droplets, or the use of fomites. Given the minimal size of the respiratory droplets, they can travel quickly to an individual's lung by inhalation [9, 10]. SARS-CoV-2 is similar in the phylogenetic sequence to SARS-CoV (79%). SARS-CoV-2, SARS-CoV, and MERS-CoV biological features, as illustrated in Table 1.
| Characteristics | SARS-CoV | MERS-CoV | SARS-CoV-2 | References |
|---|---|---|---|---|
| Outbreak | November 2002 | November 2012 | December 2019 | [11] |
| Place | Guangdong, China | Saudi Arabia | Wuhan, China | [12,13] |
| Primary reservoir | Bat, palm civet | Camel, bat | Bat | [14-16] |
| Countries affected | 26 | 27 | Over 180 | [17-19] |
| Receptor involve | ACE2 receptor | ACE2 receptor | DPP4 receptor | [20,21] |
| Diagnosis | RT-PCR, rRT-PCR |
RT-LAMP | RT-PCR, rRT-PCR |
[22-24] |
2. CORONAVIRUS TRANSMISSION
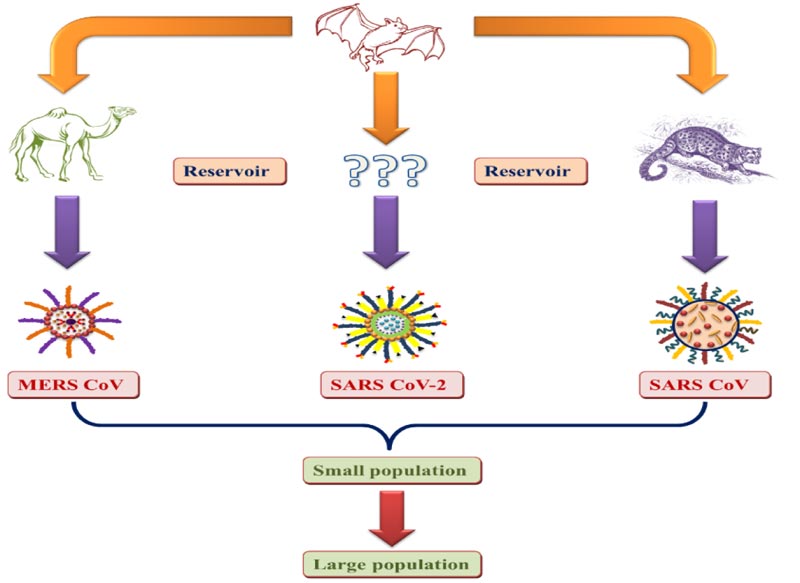
The pandemic with an unknown etiology arose from the Chinese seafood industry for the first time. The source and transmission of the virus must be determined to develop potential therapeutics. As Bat-CoV is 96.2 percent, similar to human SARS-COV-2, the bat is reported to be the primary coronavirus reservoir. The person who has a history of visiting or contacting the infected area is reported to be infected with the virus. The National Health Commission of China notified the chances of transmission between health workers. One reason for transmission was the consumption of infected animals and direct contact with primary or secondary reser- voirs. Asymptomatic infection can occur in persons with lower immune responses. The viral load found in asymptomatic patients has been found to be similar to the virus transmission capacity of symptomatic patients [25, 26]. Fig. (1) shows the transmission of coronaviruses from animals to humans.
3. THE ENTRY MECHANISM OF THE HUMAN CORONAVIRUS
The four types of structural glycoproteins are contained in coronaviruses, including Spike (S), Membrane (M), Nucleo- capsid (N), and Envelope (E). Spike glycoprotein is primarily responsible for interacting and entering the host organism. The Open-Reading Frame1 (ORF1) encodes structural proteins for all coronaviruses with unique genes [27]. Cryo-electron tomography has been shown to form an extra interior layer of the transmembrane protein in the carboxy region, thickening the viral membrane [28]. Coronaviruses are ingested according to various enzymes such as trypsin-like human Proteases in airways, cathepsins, and serine-2 Transmembrane proteases (TMPRSS2) responsible for the removal of glycoprotein. Spike (S) protein is composed of S1 and S2 subunits, responsible primarily for the binding to the host receptor and viral cell membrane fusion by forming a six-helical bundle, respectively [29, 30]. The dipeptidyl peptidase-4 (DPP-4) was reported to act as a receptor for MERS-CoV, while ACE2 was shown to be the entry receptor for SARS-CoV [21].
The SARS-CoV-2 coronavirus structure is made up of glycoprotein fusion with implicit RNA polymerase, papain-like protease, helicase, and accessory proteins. SARS-CoV-2 spiked protein is mainly attached with van der waals forces to the receptor-binding domain [31-33].
The glutamine residue 394 in SARS-CoV-2 Receptor-Binding Domain (RBD), which has a structural resemblance to 479 residues in SARS-CoV, can be identified by the essential lysine 31 on the human ACE2 receptor [34]. SARS-CoV-2 recognizes human ACE2 more prominently than SARS-CoV, responsible to increase the transmission rates from person to person [35]. N501T mutation in SARS-CoV-2 spike protein increased binding affinity to angiotensin-converting enzyme 2, causing pathogenic divergence from SARS-CoV [8].
4. GENOMIC VARIATIONS
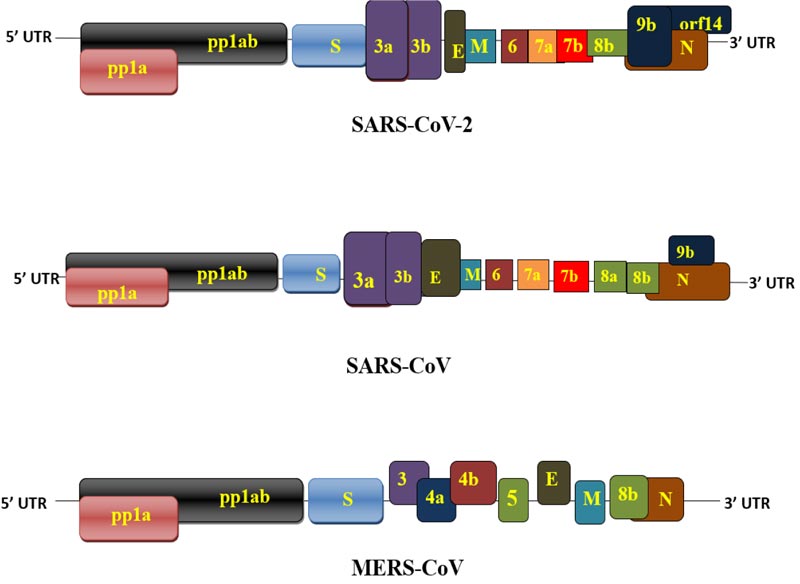
Coronavirus genome consists of approximately 26000- 32000 bases (SARS-CoV 29,712; SARS-Cov-2 ~30,000; MERS-CoV 30,119) including variability in Open Reading Frames (ORFs) [36]. The genomic sequence of SARS-CoV-2 was registered in the NCBI genome database (NC_045512.2), approximately 29.9 kb in size [37]. The genomic analysis of SARS-CoV-2 showed a similarity of 96.3%, 89% and 82% with bat CoV, SARS-like CoV, and SARS-CoV, respectively [38]. The genome of SARS-Cov-2 has 11 protein-coding genes with 12 expressed proteins. Basically, open reading frames are designed as replicase and protease (1a-1b), and major structural proteins are arranged from 5′ to 3′ order and preferred for drug targets [39]. The retrieved translated sequence of SARS-CoV-2 from GenBank showed that it encodes about 7096 long polyprotein residues with various structural and non-structural proteins [40]. The orf1ab gene in SARS-CoV-2 encodes pp1ab protein and 15 non-structural proteins (nsps), whereas the orf1a gene codes for pp1a protein and 10 non-structural proteins. The 15 nsps were categorised from nsp1 to nsp10 and nsp12 to nsp16. The orf1ab and orf1a genes are located at the 5' end and encode pp1ab and pp1a, respectively, and the 3' end of the genome contains four structural glycoproteins (S, E, M, N) and eight accessory proteins (3a, 3b, p6, 7a,7b 8b, 9b, and orf14) [31]. SARS-CoV has some differences in accessory proteins (3a, 3b, 6, 7a, 7b, 8a, 8b, and 9b) [41]. The accessory proteins help in virus transmission, initiates pathological events and produce pro-inflammatory cytokines and activate interferon signaling [38]. The genetic makeup of SARS-CoV- showed there are 380 amino acid changes from different protein to the proteins of recent SARS-CoV-2. For example, accessory proteins, S protein and N protein have 348, 27 and 5 amino acid changes, respectively [42]. The coronavirus phylogenetic tree revealed a structural similarity between SARS-CoV-2 and SARS-CoV [11, 43]. The amino acid sequence of SARS-CoV-2 is quite similar to SARS-CoV, but there are differences in 8a and 8b proteins [31]. For example, 8a protein present is in SARS-CoV but not in SARS-CoV-2; 8b protein consists of 84 amino acids in SARS-CoV, whereas amino acids in SARS-CoV; 3b protein is composed of amino acids in SARS-CoV but only 22 amino acids in SARS-CoV-2 [44].
MERS-CoV has a structural resemblance with SARS-CoV- 2. Genetically, MERS-CoV is composed of 5' cap structure, a poly (A) tail at 3' end, the rep gene consists of 16 nsps (nsp1-nsp16). At the 3' end, it consists of structural proteins (S, E, M, N) as well as 5 accessory proteins (3, 4a, 4b, 5, 8) [36, 45]. The accessory SARS-CoV-2, SARS-CoV- and MERS-CoV proteins have a certain heterogeneity, shown in Fig. (2) for their genomic variability.
5. TREATMENT STRATEGY
At the moment, coronavirus cannot be fully cured by any therapy. The primary use of antibiotics and anti-viral medicines is to relieve loads of viral RNA [46]. The combination of lopinavir and ritonavir showed clinical effectiveness against SARS-CoV but not against 2019-nCoV [47]. Remdesivir blocked, in particular 2019-nCoV replication combined with chloroquine or immune interferon [8, 48]. The results for newly-infected patients were successfully proved by isolated blood plasma from clinically treated COVID-19 patients.
5.1. Anti-viral Drugs
There are no successful anti-viral agents that can fight against COVID-19. Lopinavir is a protease inhibitor in only one in in vitro and pre-clinical studies. Anti-viral drug remdesivir has been shown to be effective against Ebola [49]. It shows efficacy against RNA viruses and can combat against RNA-dependent RNA-polymerase(RdRp) [50]. Lists of recent clinical trials of anti-viral drugs in COVID-19 patients are shown in Table 2.
|
Identification Number |
Public Title | Group 1 | Group 2 | Group 3 | Group 4 | Primary Outcome | Primary Sponsor |
|---|---|---|---|---|---|---|---|
| ChiCTR2000031734 |
Evaluation Danorevir sodium tablets combined with ritonavir in the treatment of novel Coronavirus Pneumonia (COVID-19): a randomized, open-label, controlled trial | Experimental group- Danorevir sodium tablets,/ritonavir oral (40 patients) | - | - | - | Rate of composite adverse outcomes: SpO2, PaO2/FiO2, respiratory rate | Huoshenshan Hospital |
| ChiCTR2000030472 | An open and controlled clinical study to evaluate the efficacy and safety of Ganovo combined with ritonavir in the treatment of novel coronavirus pneumonia (COVID-19) | Experimental group- Ganovo/ ritonavir oral+conventional treatment (10 patients) | Control group- Conventional treatment (10 patients) | - | - | Rate of composite adverse outcomes: SpO2, PaO2/FiO2, and respiratory rate | Shenyang Sixth People's Hospital |
| ChiCTR2000030218 | Study of Lopinavir / Ritonavir Tablets (Trade Name: Kelizhi) Combined with Xiyanping Injection for Novel Coronavirus Pneumonia (COVID-19) | Experimental group- Lopinavir/ritonavir tablets combined with Xiyanping injection (30 patients) | Control group- Keep ritonavir/ ritonavir treatment (30 patients) |
Experimental group- Lopinavir/ritonavir tablets combined with Xiyanping injection (20 patients) |
- | Pneumonia Severity Index (PSI) score | Fifth People's Hospital of Ganzhou |
| ChiCTR2000030113 | Randomized controlled trial for safety and efficacy of Favipiravir in the treatment of novel coronavirus pneumonia (COVID-19) with poorly responsive ritonavir/ritonavir | Control group-Keep ritonavir/ritonavir treatment (15 patients) | Experimental group-Favipiravir (15 patients) | - | - | Blood routine tests, Liver function examination, Renal function examination, Blood gas analysis, Chest CT Examination | The Third People's Hospital of Shenzhen |
| ChiCTR2000029603 | A Randomized, Open-Label, Multi-Centre Clinical Trial Evaluating and Comparing the Safety and Efficiency of ASC09/Ritonavir and Lopinavir/Ritonavir for Confirmed Cases of Novel Coronavirus Pneumonia (COVID-19) | Experimental group-Conventional standardized treatment and ASC09/Ritonavir (80 patients) | Control group-Conventional standardized treatment and Lopinavir/ Ritonavir (80 patients) |
- | - | The incidence of the composite adverse outcome within 14 days after admission: Defined as (one of them) SpO2<= 93% without oxygen supplementation, PaO2/FiO2 <= 300mmHg or RR <=30 breaths per minute. | The First Affiliated Hospital of Zhejiang University School of Medicine |
| ChiCTR2000029741 | Efficacy of Chloroquine and Lopinavir/ Ritonavir in mild/general novel coronavirus (COVID-19) infections: a prospective, open-label, multicenter randomized controlled clinical study | Experimental group- Chloroquine Phosphate (56 patients) | Control group- Lopinavir / Ritonavir (56 patients) | - | - | Oxygenation index during treatment; Peripheral blood cell count; C-reactive protein; procalcitonin |
The Fifth Affiliated Hospital Sun Yat-Sen University |
| ChiCTR2000030187 | Clinical study for Lopinavir and Ritonavir in the treatment of novel coronavirus pneumonia (COVID-19) | Experimental group- Lopinavir and Ritonavir Tablets (30 patients) | Control group-Routine symptomatic support treatment (30 patients) | - | - | Endotracheal intubation rate; Mortality | Jingzhou First People's Hospital |
| NCT04315948 | Multi-centre, Adaptive, Randomized Trial of the Safety and Efficacy of Treatments of COVID-19 in Hospitalized Adults |
Experimental group- Remdesivir will be administered as a 200 mg intravenous loading dose on Day 1, followed by a 100 mg once-daily intravenous upto 10 days total course (620 patients) | Experimental group- Lopinavir/ritonavir (400 lopinavir mg/100 mg ritonavir) will be administered every 12 h for 14 days in tablet form (620 patients) | Experimental group- Lopinavir/ritonavir (400 lopinavir mg/100 mg ritonavir) will be administered every 12 h for 14 days in tablet form and Interferon ß1a will be administered subcutaneously at the dose of 44 µg for a total of 3 doses in 6 days (day 1, day 3, day 6) (620 patients) | Experimental group- Hydroxychloroquine will be administered orally as a loading dose of 400 mg twice daily for one day followed by 400 mg once daily for 9 days (620 patients) | Percentage of subjects reporting each severity rating on a 7-point ordinal scale | Institut National de la Santé Et de la Recherche Médicale, France |
| NCT04321616 | The (Norwegian) NOR Solidarity Multicenter Trial on the Efficacy of Different Anti-viral Drugs in SARS-CoV-2 Infected Patients | Experimental group- Hydroxychloroquine will be given orally (in the ICU in gastrointestinal tubes) with 800 mg x 2 loading dose followed by 400 mg x 2 every day for a total of 10 days | Experimental group- Remdesivir will be given intravenously 100 mg daily for the duration of the hospitalization and up to 10 days total course. A loading dose of 200 mg at inclusion will be given | • Control group- The standard of care will be supplied to all patients not receiving a drug intervention | - | All-cause in-hospital mortality | Oslo University Hospital |
| NCT04280705 | A Multicenter, Adaptive, Randomized Blinded Controlled Trial of the Safety and Efficacy of Investigational Therapeutics for the Treatment of COVID-19 in Hospitalized Adults | Control group- Placebo Comparator: 200 mg of Remdesivir placebo administered intravenously on Day 1, followed by a 100 mg once-daily maintenance dose of Remdesivir placebo while hospitalized for up to a 10 days total course (286 patients) | Experimental group- 200 mg of Remdesivir administered intravenously on Day 1, followed by a 100 mg once-daily maintenance dose of Remdesivir while hospitalized for up to a 10 days total course (286 patients) | - | - | Time to recovery; Percentage of subjects reporting each severity rating on the 7-point ordinal scale | National Institute of Allergy and Infectious Diseases (NIAID) |
| NCT04333589 | The Mechanism, Clinical Outcome and Therapeutic Intervention of Coronavirus Disease 2019 Patients Whose Nucleic Acids Changed From Negative to Positive | Experimental group- Favipiravir group- On the 1st day, 1600mg each time, twice a day; from the 2nd to the 7th day, 600mg each time, twice a day. Oral administration, the maximum number of days taken is not more than 14 days |
- | - | - | Viral nucleic acid test negative conversion rate [ Time Frame: 5 months ] |
Peking University First Hospital |
5.2. Antiparasite Drugs
The use of chloroquine as an antiviral agent is crucial for preventing malaria, autoimmune diseases, and amoebiosis infections [51]. The studies show that intravesicular-pH controls cell function and increases the pH-endosomal required to fuse the virus into a host organism, including glycosylation trimming. Chloroquine prevents vacuole and endocytosis from moving protozoans. Chloroquine is known to be useful either as prophylaxis or as a therapeutic agent. Chloroquine enables the inflow of responsible zinc to inhibit the in-vitro function of RNA polymerase [52-54]. Hydroxychloroquine is less toxic than the analogue derivative of chloroquine. Hydroxychloro- quine was reported to show cell culture activity during the SARS-CoV epidemic. The pharmacokinetic study showed that hydroxychloroquine was found to be as effective as chloroquine in the treatment of SARS-CoV-2 due to a lack of experimental evidence [56, 57].
Ivermectin is a broad-spectrum FDA approved parasitic drug that shows activity against COVID-19 as a second-line drug. Ivermectin has a wide range of anti-viral activity against large numbers of viruses under in vitro conditions as it prevents viral replication. A single treatment with ivermectin reduced the virus to 5000 times in culture within 48 hours, but no further reduction to 72 hours. Ivermectin was known to inhibit the nuclear import of viruses and host proteins. It has been reported that the integrase protein (IN) of viruses and the importin (IMP5-007 / β1) heterodimer is responsible for IN nuclear import. As most RNA viruses rely on IMP / β1 during infection, ivermectin directly affects it and inhibits virus replication [58]. Several clinical trials to test therapeutic potency in 2019-nCOV started in different hospitals and universities. Several patient age groups were used to control adverse effects. The list of recent clinical trials of anti-parasitic drugs in COVID-19 patients is shown in Table 3.
| Identification number | Public Title | Group 1 | Group 2 | Group 3 | Group 4 | Outcome | Sponsor |
|---|---|---|---|---|---|---|---|
|
ChiCTR2000031782 |
A questionnaire investigation of hydroxychloroquine for its potential protective effect against Severe Acute Respiratory Syndromes-coronavirus-2 infection | Patients with LE and are taking hydroxychloroquine (200 patients); another group with LE does not take hydroxychloroquine (50 patients) | Patients with dermatomyositis and are taking hydroxychloroquine (200 patients); another group with dermatomyositis does not take hydroxychloroquine (100 patients) | Patients with RA and are taking hydroxychloroquine (50 patients); another group with RA are not take hydroxychloroquine (200 patients) | Patients with rosacea and are taking hydroxychloroquine (200 patients); another group with rosacea does not take hydroxychloroquine (200 patients) | Incidence of SARS-CoV-2 infection (including confirmed SARS-CoV-2 detection, but might asymptomatic) | The Second Xiangya Hospital of Central South University |
|
ChiCTR2000031204 |
A multicenter, single-blind, randomized controlled clinical trial for chloroquine phosphate in the treatment of 2019 novel coronavirus-infected pneumonia | Experimental group- Oral chloroquine phosphate tablets (150 patients) | Control group- Oral placebo group (150 patients) | - | - | Clearance time of virus RNA | Beijing you'an Hospital; Capital Medical University |
|
ChiCTR2000031174 |
Effectiveness and safety of hydroxychloroquine sulfate in the preventive treatment of novel coronavirus pneumonia (COVID-19) | Experimental group- Hydroxychloroquine (1000 patients) | Control group- Placebo (1000 patients) | - | - | COVID-19 Nucleic acid | Shanghai Public Health Clinical Center |
|
ChiCTR2000030054 |
An open randomized controlled trial for Chloroquine phosphate and Hydroxychloroquine sulfate in the treatment of mild and common novel coronavirus pneumonia (COVID-19) | Experimental group- Hydroxychloroquine sulfate 0.2g bid x 14 days a day (40 patients) |
Experimental group- The first dose of chloroquine phosphate was 1gx2 days, and the third day was 0.5gx12 days (40 patients) |
Recommended treatment plan for novel coronavirus pneumonia diagnosis and treatment plan (20 patients) |
- | Clinical recovery time | Zhongshan Hospital Affiliated to Xiamen University |
| ChiCTR2000029868 | Hydroxychloroquine treating novel coronavirus pneumonia (COVID-19): a randomized controlled, open label, multicenter trial | Experimental group- Oral hydroxychloroquine sulfate tablets (180 patients) | Conventional treatment meet the Guideline (180 patients) | - | - | Viral nucleic acid test | Ruijin Hospital; Shanghai Jiaotong University School of Medicine |
| NCT04374279 | A Phase II Trial to Promote Recovery From COVID-19 With Ivermectin or Endocrine Therapy | Experimental group- Bicalutamide 150 Mg Oral Tablet for 7 days, and standard of care | Experimental group- Ivermectin 3Mg Tab (Ivermectin 600 µg/kg (up to a maximum dose of 60mg) by mouth daily for 3 days) | - | - | Number of participants who have clinical improvement at day 7 after randomization | Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins |
| NCT04341493 | Treatment With Hydroxychloroquine vs Nitazoxanide + Hydroxychloroquine in Patients With COVID-19 With Risk Factors for Poor Outcome | Experimental group- Hydroxychloroquine 400 mg PO every 12 hours for two days and then 200 mg PO every 12 hours for four days + Nitazoxanide 500 mg PO every 6 hours for six days | Active comparator- Hydroxychloroquine 200 mg PO every 12 hours for 7 days | - | - | Mechanical ventilation requirement | Hugo Mendieta Zeron |
| NCT04363450 | Hydroxychloroquine as Primary Prophylaxis for COVID-19 in Healthcare Workers (HCQPreP) | Experimental group- Hydroxychloroquine loading dose will be given as 400mg for two doses 12 hours apart. This will then be followed by maintenance dosing of 200mg twice weekly for the remainder of the trial | Control group- Placebo: (An identical placebo will be administered on an identical dosing interval and frequency) | - | - | Incidence of symptomatic COVID-19 infection in healthcare workers | Louisiana State University Health Sciences Center in New Orleans |
| NCT04346667 | Post-Exposure Prophylaxis for Asymptomatic SARS-CoV-2 COVID-19 Patients With choloroquinE Compounds | Experimental group- Hydroxychloroquine Sulfate Regular dose (Hydroxychloroquine administered based off of in-vitro pharmacokinetics study) | Experimental group- Hydroxychloroquine Sulfate Loading Dose | Experimental group- Chloroquine administered based off of in-vitro pharmacokinetics study | Placebo (Standard of Care plus placebo) | RT-PCR negative status | Government of Punjab, Specialized Healthcare and Medical Education Department |
| NCT04342169 | Hydroxychloroquine for Outpatients With Confirmed COVID-19 | Experimental group- Hydroxychloroquine (HCQ 400mg po BID x 1 day, then 200mg po BID x 4 days) | Control group- Placebo oral tablet (Placebo to be taken on the same schedule as HCQ) | - | - | Duration of viral shedding | University of Utah |
| NCT04341441 | Will Hydroxychloroquine Impede or Prevent COVID-19: WHIP COVID-19 Study | Experimental group- Hydroxychloroquine - Daily Dosing (The daily hydroxychloroquine treatment arm will receive a 200 mg oral dose daily following day 1 dose of 400 mg orally once) | Experimental group- Hydroxychloroquine - Weekly Dosing (The once weekly randomized treatment arm will receive the proposed dose of hydroxychloroquine for prophylaxis of malaria is 6.5 mg/kg per dose (maximum of 400 mg per dose) administered orally weekly on the same day of each week) | Placebo oral tablet (Participants randomized to this arm will be provided with daily dosing of oral placebo to have the patients take 2 pills a day) | Non-Randomized Active Comparator | Reduction in the number of COVID-19 infections in healthcare workers | Henry Ford Health System |
| NCT04392427 | Effect of a Combination of Nitazoxanide, Ribavirin and Ivermectin Plus Zinc Supplement on the Clearance of COVID-19: a Pilot Sequential Clinical Trial | Experimental group- Will receive a combination of Nitazoxanide, Ribavirin and Ivermectin or a duration of seven days (100 patients) | Control group- will not receive anything | - | - | PCR FOR COVID-19 will be done on serial visits till turn to negative, first after 5 day then serial every 48 hours till become negative for two consecutive samples. | Mansoura University |
| NCT04390022 | Pilot Study to Evaluate the Potential of Ivermectin to Reduce COVID-19 Transmission | Experimental group- Participants on this arm will receive a single, oral dose of Ivermectin 400 mcg/kg at the enrolment visit | Control group- Participants on the arm will receive a single, oral dose of placebo tablets at the enrolment visit | - | - | Proportion of patients with a positive SARS-CoV-2 PCR from a nasopharyngeal swab at day 7 post-treatment | Clinica University of Navarra, University of Navarra |
5.3. Corticosteroids
Corticosteroids are a group of steroid hormones that regulate various physiological processes. The protective effect of steroids in COVID-19 patients was seen in various clinical studies. Several studies have demonstrated the effectiveness of corticosteroids in alleviating adverse immune system reactions. A lab study of dexamethasone infected pigs showed that one or two doses of corticosteroids could reduce cytokine expression [59, 60]. List of recent corticosteroid clinical trials in COVID- 19 patients is shown in Table 4.
5.4. Antibodies
Monoclonal antibodies are mainly targeted at the spike glycoprotein virus that invades host organisms. There are two functional subunits of spike protein (S1 and S2), in which S1 is used to attach cells, and S2 is capable of fusing into the cells. Monoclonal antibodies can only be monovalent, and only one antigen can be identified at the same time. Antibodies neutra- lizing coronavirus are frequently targeted at and make incompetent S1 binding receptor domains [62, 63]. Some antibodies identify various epitopes in the domain of receptor bindings, such as SARS-CoV neutralizing the virus compe- tency antibodies CR 3014 and CR 3022. Table 5 shows the list of recent clinical tests of antibodies in COVID-19 patients.
| Identification Number | Public Title | Group 1 | Group 2 | Group 3 | Group 4 | Outcome | Sponsor |
|---|---|---|---|---|---|---|---|
| NCT04355637 | Inhaled Corticosteroid Treatment of COVID19 Patients With Pneumonia | Control group- Patients receiving standard of care to treat their pneumonia | Experimental group- Patients receiving standard of care to treat their pneumonia + inhaled budesonide | - | - | Proportion of patients in both arms fulfilling the criteria for treatment failure | Sara Varea |
| NCT04344288 | Corticosteroids During COVID-19 viral Pneumonia Related to SARS-Cov-2 Infection | Experimental group- Oral Prednisone during 10 days (0.75 mg/kg/day during 5 days then 20 mg/day during 5 more days) | Control group-Standard of care | - | - | Number of patients with a theoretical respiratory indication for transfer to intensive care unit evaluated by a SpO2 <90% stabilized at rest and under not more than 5 L / min of supplemental oxygen using medium concentration mask | Hospices Civils de Lyon |
| NCT04355247 | Prophylactic Corticosteroids to Prevent COVID-19 Cytokine Storm | Experimental group- Methylprednisolone 80 mg/mL Injectable Suspension will be given daily x 5 days starting upon day 1 of admission to hospital. | - | - | - | Clinical complete response criteria; Clinical Partial Response criteria | Auxilio Mutuo Cancer Center |
| NCT04343729 | Methylprednisolone in the Treatment of Patients With Signs of Severe Acute Respiratory Syndrome in COVID-19 | Experimental group- 0.5mg/kg injectable methylprednisolone sodium succinate, twice daily, for 5 days. | Control group- Saline solution, twice daily, for 5 days. Injectable | - | - | Mortality rate at day 28 | Fundação de Medicina Tropical Dr. Heitor Vieira Dourado |
| ChiCTR2000030481 | The clinical value of corticosteroid therapy timing in the treatment of novel coronavirus pneumonia (COVID-19): a prospective randomized controlled trial | Experimental group- Early corticosteroid intervention group (75 patients) | Experimental group- Middle-late corticosteroid intervention group (75 patients) | Control group- No corticosteroid (50 patients) | - | The time of duration of COVID-19 nucleic acid RT-PCR test results of respiratory specimens (such as throat swabs) or blood specimens change to negative. | Zhongnan Hospital of Wuhan University |
| Identification Number | Public Title | Group 1 | Group 2 | Group 3 | Group 4 | Outcome | Sponsor |
|---|---|---|---|---|---|---|---|
| NCT04346589 | Convalescent Antibodies Infusion in Critically Ill COVID 19 Patients | Experimental group- Antibodies (immunoglobulins) infusion- Biological: Anti-coronavirus Antibodies (immunoglobulins) obtained with DFPP from convalescent patients |
- | - | - | Number of mechanical ventilation days | A.O. Ospedale Papa Giovanni XXIII |
| NCT04341116 | Study of TJ003234 (Anti-GM-CSF Monoclonal Antibody) in Subjects With Severe Coronavirus Disease 2019 (COVID-19) | Experimental group- TJ003234 (3mg/kg): patients receive a single infusion | Experimental group- TJ003234 (6mg/kg): patients receive a single infusion | Control group- Placebo: patients receive a single infusion | - | Proportion (%) of subjects experiencing deterioration in clinical status | I-Mab Biopharma Co. Ltd. |
| NCT04351152 | A Phase 3 Randomized, Placebo-Controlled Study of Lenzilumab in Hospitalized Patients With COVID-19 Pneumonia | Experimental group- Lenzilumab IV infusion plus Standard of Care | Control group- IV infusion of saline plus Standard of Care | - | - | Incidence of invasive mechanical ventilation (IMV) and/or Mortality | Humanigen, Inc. |
| ChiCTR2000030703 | A randomized, blinded, controlled, multicenter clinical trial to evaluate the efficacy and safety of Ixekizumab combined with conventional antiviral drugs in patients with novel coronavirus pneumonia (COVID-19) | Experimental group- Ixekizumab and antiviral therapy (20 patients) | control group- antiviral therapy (20 patients) | - | - | Lung CT; Lung function; Arterial blood gas analysis | Xiangya Hospital of Central South University |
|
Identification Number |
Public Title | Group 1 | Group 2 | Group 3 | Group 4 | Outcome | Sponsor |
|---|---|---|---|---|---|---|---|
| ChiCTR2000030039 | Clinical study for infusing convalescent plasma to treat patients with new coronavirus pneumonia (COVID-19) | Experimental group-Conventional therapy with Infusion of convalescent plasma: 200-500ml, two infusions are recommended (30 patients) | Control group -Conventional therapy (60 patients) | - | - | SARS-CoV-2 DNA; SARS-CoV-2 antibody levels | Affiliated Hospital of Xuzhou Medical University |
| ChiCTR2000030929 | A randomized, double-blind, parallel-controlled trial to evaluate the efficacy and safety of anti-SARS-CoV-2 virus inactivated plasma in the treatment of severe novel coronavirus pneumonia (COVID-19) | Experimental group-Anti-SARS-CoV-2 virus inactivated plasma (30 patients) | Control group- Ordinary plasma (30 patients) | - | - | Improvement of clinical symptoms | Renmin Hospital of Wuhan University |
| ChiCTR2000030702 | Convalescent plasma for the treatment of common COVID-19: a prospective randomized controlled trial | Experimental group-Conventional treatment and convalescent plasma therapy (25 patients) | Control group- Conventional treatment (25 patients) | - | - | Time to clinical recovery after randomization | China-Japan friendship hospital |
| NCT04358783 | Convalescent Plasma Compared to the Best Available Therapy for the Treatment of SARS-CoV-2 Pneumonia | Experimental group-Convalescent Plasma from cured COVID-19 patients and supportive management depending on individual needs | Experimental group- Receive supportive management depending on individual needs | - | - | - | Hospital Universitario Dr. Jose E. Gonzalez |
| NCT04357106 | COPLA Study: Treatment of Severe Forms of Coronavirus infection With Convalescent Plasma | Experimental group- 200 ml of convalescent Plasma, single dose | - | - | - | Lung injury(PaO2/FiO2 relation); Overall survival | Centro de Hematología y Medicina Interna |
| NCT04360486 | Treatment Of CORONAVIRUS DISEASE 2019 (COVID-19) With Anti-Sars-CoV-2 Convalescent Plasma (ASCoV2CP) |
Anti-Sars-CoV-2 Convalescent Plasma- Fresh frozen plasma, plasma Frozen for 24 hours (PF-24), or liquid plasma |
- | - | - | - | U.S. Army Medical Research and Development Command |
| NCT04377672 | Safety and Pharmacokinetics of Human Convalescent Plasma in High Risk Children Exposed or Infected With SARS-CoV-2 | Experimental group- Anti-SARS-CoV-2 Human Convalescent Plasma (1-2 units (200-250 mL per unit) of plasma with anti-SARS-CoV-19 titers of ≥1:320). The total volume (mL) infused will be based on weight (5 mL/kg) with a maximum volume of 500 mL | - | - | - | Safety of treatment with high-titer anti-SARS-CoV-2 plasma as assessed by adverse events | Johns Hopkins University |
| NCT04364737 | Convalescent Plasma to Limit COVID-19 Complications in Hospitalized Patients | Experimental group- SARS-CoV-2 convalescent plasma (1-2 units; ~250-500 mL) | Control group- Lactated ringer's solution or sterile saline solution | - | - | Percentage of subjects reporting each severity rating on WHO ordinal scale for clinical improvement | NYU Langone Health |
| Identification Number | Public Title | Group 1 | Group 2 | Group 3 | Group 4 | Outcome | Sponsor |
|---|---|---|---|---|---|---|---|
| ChiCTR2000031781 | A randomized, double-blinded, placebo-controlled phase II clinical trial for Recombinant Novel Coronavirus (2019-nCOV) Vaccine (Adenovirus Vector) in healthy adults aged above 18 years | Experimental group- Middle dose (1E11vp) (250 patients) |
Experimental group- Low dose (5E10vp) (125 patients) | Control group- Placebo (125 patients) |
- | Adverse reactions 0-14 days post vaccination; Anti-S antibody IgG titer on day 28 post vaccination; Anti-SARS-CoV-2 neutralizing antibody titer on day 28 post vaccination | Jiangsu Provincial Center for Disease Control and Prevention |
| NCT04276896 | Phase I/II Multicenter Trial of Lentiviral Minigene vaccine (LV-SMENP) of COVID-19 Coronavirus | Experimental group- Injection and infusion of LV-SMENP-DC vaccine and antigen-specific CTLs (Patients will receive approximately 5x10^6 LV-DC vaccine and 1x10^8 CTLs via sub-cutaneous injections and iv infusions, respectively. | - | - | - | Clinical improvement based on the 7-point scale; Lower Murray lung injury score | Shenzhen Geno-Immune Medical Institute |
| NCT04348370 | Bacillus Calmette-Guerin vaccination as Defense Against SARS-CoV-2: A Randomized Controlled Trial to Protect Health Care Workers by Enhanced Trained Immune Responses | Experimental group-BCG vaccine (A single dose will consist of 0.1 mL (~2x10^5 CFU) will be administered by slow intradermal injection using a 25 gauge/ 0.5 mm syringe in the deltoid area.) | Control group- Vaccine (A single dose will consist of 0.1 mL saline) | - | - | Incidence of COVID 19 Infection [Time Frame: 6 months] | Texas A&M University |
| NCT04361552 | Tociluzumab for Cytokine Release Syndrome With SARS-CoV-2: An Open-Labeled, Randomized Phase 3 Trial | Experimental group- Arm I: Patients receive tocilizumab IV every 12 hours for up to 3 doses in the absence of disease progression or unacceptable toxicity. Patients also receive standard of care | Active Comparator- Arm II: Patients receive standard of care | - | - | 7-day length of invasive mechanical ventilation (MV) ; 30-day mortality rate | Emory University |
| NCT04359667 | Serum IL-6 and Soluble IL-6 Receptor in Severe COVID-19 Pneumonia Treated With Tocilizumab | Experimental group- Tocilizumab 20 MG/ML Intravenous Solution [ACTEMRA] (1 - 8 mg per kg of body weight once, maximal 800 mg per dose and also standard of care treatment) | - | - | - | Evaluate the role of laboratory markers as predictors of survival | University Hospital for Infectious Diseases, Croatia |
| NCT04283461 | Phase I, Open-Label, Dose-Ranging Study of the Safety and Immunogenicity of 2019-nCoV Vaccine (mRNA-1273) in Healthy Adults | Experimental three different groups receive 25,100,250 mcg of mRNA-1273, respectively, administered through IM between 18-55 years age range | Experimental three different groups receive 25,100,250 mcg of mRNA-1273, respectively, administered through IM between 56-70 years age range | Experimental three different groups receive 25,100,250 mcg of mRNA-1273, respectively, administered through IM with age 71 years or older | - | Frequency of solicited local reactogenicity adverse events (AEs) [ Time Frame: Through 7 days post-vaccination ]; Frequency of any medically-attended adverse events (MAAEs) [ Time Frame: Day 1 to Day 394 ] |
National Institute of Allergy and Infectious Diseases (NIAID) |
5.5. Transfusion of Convalescent Plasma
The administration of convalescent plasma to SARS-CoV- 2 infected patients shows recovery from the virus’s etiology and pooled mortality rates as significantly decreased compared with or without placebo [64-66]. The health commissions of various backgrounds have asked recovered patients for donating their blood. Patients who received convalescent plasma reported a rapid recovery within 14 days, compared with other patients during the SARS CoV outbreak [67]. Table 6 shows the list of recent clinical trials of plasma therapies in patients with COVID-19.
5.6. Vaccines
The already revealed interaction among host receptors with coronavirus allows researchers to find a cure for nCoV 2019. In recent centuries, vaccination in severe diseases has been a significant defensive function. A clinic trial of six vaccines was carried out to test the efficacy of these vaccines, including mRNA1273(NCT042834461), S-protein adenoviral type 5 (NCT04313127), Chimpanzee adenoviral vector ChAdOx1 (NCT04324606), S-protein plasm encoding (NCT04336410), Lentiviral DCs modified (NCT04276896) and artificial antigen cells modified with lentiviral vector expression. The clinical trial without pre-clinical studies was concluded in a very short period because of the high and safe therapy potential of mRNA1279-COVID-19 (NCT04283461) encapsulated nano- particles [68]. The safety profile of the mRNA vaccine is outstanding and has excellent immunological properties. mRNA vaccines are mostly induced by cellular and humoral immunity [69]. A list of recent clinical trials of vaccines in COVID-19 patients shown in Table 7 can be a game-changer for vaccine technologies.
CONCLUSION: FUTURE ASPECTS
Over the past two decades, coronavirus has shown worldwide health concerns. The disease is likely linked to hematological and respiratory problems. The spike protein of SARS-CoV-2 is more likely to reach the host compared to SARS-CoV’s spike protein, which means the transfer rate in the SARS-CoV-2 is high. Asymptomatic patients also have a high transmission rate. The host immune response must be improved to fight against coronavirus. The intermediate reservoir of nCoV-2019 is still challenging for researchers. Coronaviruses are significantly attached to the ACE2 receptor of host organisms. The open reading frame of coronaviruses is responsible to distinguish between SARS-CoV-2 and SARS-CoV. Various clinical studies have started to identify potential therapies for eradicating this pandemic, and, until now, no effective nCoV-2019 drugs or vaccines are available. All drugs are based on the experience of SARS, MERS, and other strained viruses. Running clinical trials must be focused on quality data that can be used in possible prevention and treatments. In addition to medicines, techniques for respiratory support and modulation of immune status are highly required. Global resources with reasonable scientific justification are available for the planning of clinical trials. In recent clinical trials, the repurposing and repositioning of certain drugs have been processed. The repurposing of medicines has some barriers while repositioning clinical studies facilitate the discovery of new drugs. By this year, the way to find COVID-19 solutions should be through global cooperation with different clinical trial hospitals with a large number of patients. More work is required to find out exactly how this coronavirus is being approached.
LIST OF ABBREVIATIONS
| SARS-CoV | = Severe Acute Respiratory Syndrome Coronavirus |
| MERS-CoV | = Middle East Respiratory Syndromecoronavirus |
| nCoV | = novel Coronavirus |
| S | = Spike protein |
| E | = Envelope protein |
| M | = Membrane protein |
| N | = Nucleocapsid protein |
| ACE2 | = Angiotensin-Converting Enzyme |
| ORFs | = Open Reading Frames |
| UTR | = Untranslated region |
| RdRp | = RNA dependent RNA polymerase |
| nsp | = Non-structural protein |
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The author confirm that this article content has no conflicts of interest.
ACKNOWLEDGEMENTS
The authors express their gratitude to Chairman, Mr. Parveen Garg and Director, Dr.G.D.Gupta, ISF College of Pharmacy, Moga (Punjab), India, for their excellent vision and support.
